Strategic Guide to Minimizing Primer-Dimer Formation in PCR: From Foundational Concepts to Advanced Applications
This comprehensive article provides researchers, scientists, and drug development professionals with a systematic framework for understanding, preventing, and troubleshooting primer-dimer formation in PCR.
Strategic Guide to Minimizing Primer-Dimer Formation in PCR: From Foundational Concepts to Advanced Applications
Abstract
This comprehensive article provides researchers, scientists, and drug development professionals with a systematic framework for understanding, preventing, and troubleshooting primer-dimer formation in PCR. Covering foundational principles to advanced validation techniques, the content explores the molecular mechanisms behind primer-dimer artifacts, presents optimized primer design strategies and cycling conditions, details practical troubleshooting protocols for common laboratory scenarios, and examines rigorous validation approaches for method transfer and comparative platform analysis. The integrated guidance enables professionals to enhance assay specificity, improve quantification accuracy, and ensure reliable results across diverse molecular applications from basic research to clinical diagnostics.
Understanding Primer-Dimer Formation: Mechanisms, Impacts, and Common Causes
What is a primer dimer?
A primer dimer (PD) is a small, unintended by-product of the polymerase chain reaction (PCR) [1]. It is a short DNA fragment that is formed and amplified when PCR primers anneal to each other via complementary base pairs, rather than to the intended target DNA template [2] [3]. The amplification of these artifacts consumes PCR reagents, which can compete with and inhibit the amplification of the target DNA sequence [1].
There are two principal types of primer dimers:
- Homodimers: Formed when two identical primers (e.g., two forward primers) bind to each other [4].
- Heterodimers: Formed when two different primers (e.g., a forward and a reverse primer) bind to each other [3] [4].
How do primer dimers form?
The formation and amplification of a primer dimer is a step-wise process that can occur when primers have complementary sequences, particularly at their 3' ends [1] [5]. The mechanism is illustrated below and can be described in three key steps:
This process is often initiated at low temperatures, such as when the PCR reaction is being prepared at room temperature, because some DNA polymerases retain enzymatic activity under these conditions [1] [4].
What are the consequences of primer dimer formation?
The formation of primer dimers has several negative impacts on PCR experiments:
- Consumption of Resources: Primer dimers compete for primers, nucleotides, and DNA polymerase, reducing the efficiency and sensitivity of target DNA amplification [1] [6].
- False Positives in qPCR: In quantitative PCR (qPCR) using intercalating dyes, primer dimers generate a fluorescence signal that can be mistaken for target amplification, leading to inaccurate quantification [1] [4].
- Interference in Analysis: On an agarose gel, primer dimers appear as a diffuse smear or band between 30-100 base pairs, which can obscure the interpretation of results [2] [4]. They can also cause background noise in DNA sequencing [4].
How can I detect primer dimers?
| Method | How It Works | Characteristic of Primer Dimers |
|---|---|---|
| Agarose Gel Electrophoresis [2] [4] | PCR products are separated by size on a gel. | A smeary band or fuzzy smear at 30-50 bp (typically below 100 bp). |
| Melting Curve Analysis (for qPCR) [1] [4] | After amplification, temperature is gradually increased while fluorescence is measured. PDs melt at a lower temperature than the specific target amplicon, producing a distinct peak. | |
| No-Template Control (NTC) [2] | A control reaction is run without any template DNA. | Amplification in the NTC indicates primer dimer formation or contamination, as there is no target to amplify. |
How can I prevent primer dimer formation?
Preventing primer dimers involves strategies in primer design, reaction optimization, and the use of specialized enzymes. The following diagram summarizes the main troubleshooting pathways.
The following table provides detailed methodologies and experimental protocols for the key prevention strategies.
| Strategy | Experimental Protocol & Key Parameters | Rationale |
|---|---|---|
| Optimal Primer Design [7] [8] | Protocol: Use software (e.g., Primer3) to design primers. Manually check for 3' end complementarity. Parameters:• Length: 18-24 nucleotides.• Tm: 52-58°C for both primers, with a difference < 5°C.• GC Content: 40-60%.• 3' End: Avoid runs of 3 or more G/C bases and significant complementarity between primers. | Minimizes the chance of primers annealing to themselves or each other instead of the template [5] [9]. |
| Wet-Lab Optimization [2] [5] [9] | Protocol: Set up a series of test reactions to titrate the key parameters listed. Parameters:• Annealing Temperature: Use a gradient PCR to test temperatures 3-5°C above the calculated Tm.• Primer Concentration: Test a range from 0.1-1.0 µM; often 0.2-0.5 µM is sufficient.• Hot-Start Polymerase: Use a hot-start enzyme to prevent activity at low temperatures. | Increases stringency to favor specific primer-template binding and reduces low-temperature artifacts [1] [2]. |
| Advanced Reagents [1] [6] | Protocol: Substitute standard primers with modified versions. Reagents:• SAMRS (Self-Avoiding Molecular Recognition Systems): Nucleotide analogues that bind to natural DNA but not to other SAMRS, preventing primer-primer interactions.• rhPCR (RNase H-dependent PCR): Use primers with a blocking group that is removed only at high temperature by a thermostable RNase HII. | Chemically prevents extension from mis-annealed primers, offering a high level of specificity, especially in multiplex assays [6]. |
Research Reagent Solutions
The following table lists key reagents used to prevent primer dimer formation, along with their functions in experimental protocols.
| Reagent | Function in Prevention of Primer Dimers |
|---|---|
| Hot-Start DNA Polymerase [1] [2] | Enzyme inactive during reaction setup; activated at high temperature (e.g., 95°C) to prevent extension from primers that annealed at low temperatures. |
| SAMRS Nucleotides [1] [6] | Synthetic nucleotide analogues that incorporate into primers; bind to natural DNA but not to other SAMRS, thereby avoiding primer-primer interactions. |
| Blocked-Cleavable Primers (rhPCR) [1] | Primers with a chemical block at the 3' end; the block is removed only when the primer is perfectly matched to its template, preventing extension from dimerized primers. |
| DMSO [9] | Additive that reduces secondary structure and can lower the melting temperature (Tm), helping to improve specificity and reduce mis-priming under some conditions. |
| Magnesium Chloride (MgCl₂) [9] | Essential cofactor for DNA polymerase; optimizing its concentration (typically 1.5-2.0 mM) is critical, as excess Mg²⁺ can promote non-specific priming and dimer formation. |
Primer dimers are short, unintended DNA artifacts that form when PCR primers anneal to each other instead of binding to the intended target DNA template. These structures represent a significant challenge in molecular biology, consuming reaction resources and compromising data integrity. This technical guide examines the practical consequences of primer dimer formation and provides researchers with proven methodologies for identification, prevention, and troubleshooting.
FAQ: Understanding Primer Dimers
What are primer dimers and how do they form?
Primer dimers are small, unintended DNA fragments that can form during polymerase chain reaction (PCR) when primers anneal to each other rather than to the target template DNA [2]. They typically appear as fuzzy smears below 100 bp on gel electrophoresis [2].
Formation occurs through two primary mechanisms:
- Self-dimerization: A single primer contains regions complementary to itself, creating a free 3' end that DNA polymerase can extend [2].
- Cross-dimerization: Two different primers have complementary regions that allow them to bind together, creating extendable 3' ends [2] [10].
The greatest opportunity for primer dimer formation occurs before PCR cycling begins, when reaction components are mixed at room temperature [2] [10].
What are the specific consequences of primer dimers in PCR experiments?
Primer dimers impact PCR efficiency and data accuracy through multiple mechanisms:
Resource Depletion: Primer dimers consume primers, dNTPs, and polymerase activity that would otherwise amplify the target sequence [6] [10]. This resource competition becomes particularly problematic when target molecules are scarce [6].
Reduced Amplification Efficiency: By sequestering reaction components, primer dimers decrease the yield of desired PCR products [11] [10]. This can manifest as increased Ct values in qPCR experiments [10].
Data Interpretation Problems:
- False Positives: In SYBR Green-based qPCR, primer dimers can generate amplification signals in no-template controls, potentially leading to false positive interpretations [10].
- False Negatives: At low target concentrations, primer dimer formation can prevent target amplification altogether, resulting in false negatives [10].
Table 1: Quantitative Impact of Primer Dimers on PCR Efficiency
| Parameter Affected | Impact Level | Experimental Consequence |
|---|---|---|
| Primer availability | High | Reduced target amplification efficiency |
| dNTP consumption | Moderate to High | Resource depletion for target amplification |
| Polymerase activity | Moderate | Enzyme diverted to non-productive synthesis |
| Detection sensitivity | Variable | Increased Ct values or failed amplification |
| Signal specificity | High | Background noise in fluorescent detection |
How can I identify primer dimers in my experiments?
Gel Electrophoresis Identification:
- Short length: Primer dimers typically migrate below 100 bp, beneath the last band of standard DNA ladders [2].
- Smeary appearance: They appear as fuzzy, diffuse bands rather than well-defined discrete bands [2].
- Extended electrophoresis: Running the gel longer helps separate primer dimers from desired PCR products [2].
Control Reactions:
- No-Template Control (NTC): Including an NTC reaction is essential for identification. Since primer dimers form without template DNA, they will appear as the primary amplification product in NTC reactions [2].
- SYBR Green Detection: In qPCR with intercalating dyes, primer dimers produce characteristic amplification curves in NTC wells [10].
Troubleshooting Guide: Preventing and Resolving Primer Dimer Formation
Primer Design Optimization
Effective primer design represents the most robust approach to minimizing primer dimer formation:
Table 2: Optimal Primer Design Parameters to Minimize Dimer Formation
| Design Parameter | Optimal Value | Rationale |
|---|---|---|
| Primer length | 18-30 bases [12] | Balances specificity and binding efficiency |
| GC content | 40-60% [12] | Prevents overly stable non-specific interactions |
| 3'-end complementarity | ≤3 contiguous bases [12] [8] | Minimizes primer-primer annealing |
| Self-complementarity | ≤3 contiguous bases [12] | Reduces hairpin formation and self-dimerization |
| Tm difference between primers | ≤5°C [12] | Ensures balanced annealing efficiency |
| Melting temperature (Tm) | 55-72°C [12] | Allows sufficiently stringent annealing |
Advanced Design Strategies:
- Computational Tools: Utilize primer design software (Primer3, Oligo) that incorporates thermodynamic "nearest neighbor" calculations to predict primer interactions [12].
- SAMRS Technology: Incorporation of Self-Avoiding Molecular Recognition Systems (SAMRS) nucleotides creates primers that pair with natural DNA but not with other SAMRS components, significantly reducing primer-dimer potential [6].
- 3'-End Stability: Avoid GC-rich sequences at the 3' end, as they increase dimerization potential [5].
Reaction Condition Optimization
Thermal Cycling Parameters:
- Annealing Temperature: Increase annealing temperature incrementally (1-2°C steps) to enhance specificity. Use gradient PCR to identify optimal temperature [2] [13] [5].
- Hot-Start Activation: Utilize hot-start DNA polymerases that remain inactive until a high-temperature activation step (typically 94-95°C) [2] [13]. This prevents enzymatic activity during reaction setup when primer dimer formation is most likely [2] [10].
- Reduced Cycle Number: Limit PCR cycles to 30-35 when possible, as excessive cycling promotes primer dimer accumulation [5].
Reaction Composition:
- Primer Concentration: Optimize primer concentration (typically 0.1-1 μM) using concentration gradients. High primer concentrations promote dimer formation [2] [13] [8].
- Mg²⁺ Concentration: Excessive Mg²⁺ can promote non-specific amplification. Optimize concentration based on polymerase requirements [13] [5].
- Template Quality: Ensure template DNA is pure and intact to facilitate efficient primer binding [13].
Experimental Protocol: Systematic Optimization for Primer-Dimer Reduction
Materials and Reagents:
- Hot-start DNA polymerase [2] [13]
- HPLC-purified primers [5]
- Gradient thermal cycler
- Gel electrophoresis equipment
- SYBR Green master mix (for qPCR optimization)
Procedure:
- Initial Primer Screening:
Gradient PCR Optimization:
- Set up reactions with a temperature gradient spanning 5-10°C below and above the calculated Tm.
- Include no-template controls for each primer set.
- Analyze results by gel electrophoresis to identify temperature with specific amplification and minimal dimer formation.
Primer Concentration Titration:
- Test primer concentrations from 0.1-1 μM in 0.2 μM increments.
- Maintain constant template and reaction conditions.
- Select the lowest concentration that provides robust target amplification.
Validation with No-Template Controls:
- Perform final reactions with optimized conditions including NTC.
- For qPCR, ensure NTC shows no amplification or late Ct values (>10 cycles after lowest sample Ct) [10].
Diagram 1: PCR Optimization Workflow (46 characters)
Research Reagent Solutions
Table 3: Essential Reagents for Primer Dimer Prevention
| Reagent Category | Specific Examples | Function & Rationale |
|---|---|---|
| Hot-Start Polymerases | Hot-start Taq, Bst 2.0 WarmStart [2] [14] | Prevents enzymatic activity during reaction setup, reducing pre-cycling dimer formation |
| Primer Design Tools | Primer3, Primer Express, Oligo [12] | Computational prediction of primer interactions and dimer potential |
| Modified Nucleotides | SAMRS components [6] | Create primers that avoid self-annealing while maintaining target binding |
| PCR Additives | Betaine, DMSO [14] | Reduces secondary structure and improves specificity in complex templates |
| Purification Methods | HPLC purification [5] | Ensures primer quality and removes truncated sequences that promote dimers |
| Specialized Buffers | Isothermal amplification buffer with Mg++ optimization [14] | Provides optimal ionic conditions for specific amplification |
Advanced Applications and Considerations
Multiplex PCR Challenges
In multiplex PCR reactions containing multiple primer sets, the probability of primer dimer formation increases significantly due to higher total primer concentrations and greater opportunity for inter-primer complementarity [10] [14]. Special considerations include:
- Enhanced Computational Analysis: Use tools capable of analyzing multiple primer interactions simultaneously [10].
- Staged Primer Addition: Implement protocols where primers are added at different concentrations or after an initial hot-start activation [6].
- Empirical Validation: Thoroughly test all primer combinations with appropriate controls [14].
Isothermal Amplification Methods
Loop-mediated isothermal amplification (LAMP) utilizes 4-6 primers targeting 6-8 regions, creating substantial potential for primer dimers and hairpin structures [14]. Inner primers (FIP/BIP) are particularly problematic due to their length (typically 40-45 bases) [14]. Mitigation strategies include:
- Thermodynamic Analysis: Calculate stability of potential secondary structures using nearest-neighbor models [14].
- Primer Modification: Strategic nucleotide substitutions to disrupt stable hairpins while maintaining target binding [14].
- QUASR Detection: Using quenched primers to reduce background signal from primer dimers [14].
Primer dimers represent a significant challenge in PCR that directly impacts experimental efficiency and data accuracy. Through strategic primer design, reaction optimization, and appropriate control implementation, researchers can effectively minimize primer dimer formation. The approaches outlined in this guide provide a systematic framework for troubleshooting and preventing primer dimers across various PCR applications, ensuring reliable and interpretable results in molecular biology research.
FAQs on Primer-Dimer Formation
1. What is a primer dimer and how does it affect my PCR reaction? A primer dimer is a small, unintended DNA fragment that forms when PCR primers anneal to each other instead of binding to the intended target DNA template. This occurs through self-dimerization (a single primer binding to itself) or cross-dimerization (forward and reverse primers binding to each other) [2]. Primer dimers consume reaction reagents (primers, polymerase, dNTPs), compete with the desired amplification product for resources, and can lead to reduced target yield, false positives, or inaccurate quantification, especially in sensitive applications like qPCR [11] [6].
2. Can primer dimers form even if my primer sequences are well-designed? Yes. While careful primer design is the first line of defense, primer dimers can still form due to suboptimal reaction conditions. Factors such as low annealing temperatures, high primer concentrations, or the presence of DNA polymerase activity during reaction setup at room temperature can all promote primer-dimer formation, even with well-designed primers [2] [13]. Using a hot-start polymerase is a key strategy to prevent dimers that form during setup [2].
3. How can I confirm that a band on my gel is a primer dimer? Primer dimers have two telltale characteristics on an agarose gel: they are short (typically below 100 bp) and have a fuzzy, smeary appearance rather than a sharp, well-defined band [2]. Running a No-Template Control (NTC) is a definitive test; since primer dimers do not require a DNA template to form, they will appear as the primary product in an NTC lane [2].
4. My template has high GC content. How does this contribute to amplification problems? GC-rich sequences (over 65%) form strong secondary structures and are more stable, making them difficult to denature completely during the PCR cycle. This can reduce efficiency and promote non-specific priming, including primer-dimer formation, as primers may find it harder to access their intended binding sites [15] [13]. The use of PCR additives like DMSO or betaine can help denature these stable structures and improve amplification specificity [15].
Troubleshooting Guide
This guide addresses common issues related to template quality, reaction setup, and thermal cycling.
Table: Troubleshooting Common PCR Problems
| Symptom | Possible Cause | Recommended Solution |
|---|---|---|
| No amplification product | Degraded or insufficient template DNA [13] | Check DNA integrity by gel electrophoresis; increase amount of input DNA [13] [16]. |
| Presence of PCR inhibitors (e.g., phenol, EDTA) [15] [13] | Re-purify template DNA; dilute template to reduce inhibitor concentration; use polymerases tolerant to inhibitors [15]. | |
| Smear or multiple non-specific bands | Annealing temperature too low [15] [13] | Increase annealing temperature in 1-2°C increments; use a gradient PCR cycler for optimization [15]. |
| Excess Mg2+ concentration [13] [16] | Titrate Mg2+ concentration, typically between 1.5-5.0 mM, to find the optimal level [15] [16]. | |
| Primer-dimer formation | High primer concentration [2] [13] | Lower primer concentration (standard range is 0.1-1.0 µM) [13] [16]. |
| Non-hot-start DNA polymerase [2] [13] | Switch to a hot-start DNA polymerase to prevent activity during reaction setup [2]. | |
| Low annealing temperature [2] [15] | Increase the annealing temperature to improve stringency [2]. | |
| Low yield of desired product | Suboptimal extension time or temperature [13] | Increase extension time for longer amplicons; ensure extension temperature is optimal for the polymerase (typically 68-72°C) [13]. |
| Poor primer design [15] [13] | Redesign primers to avoid self-complementarity and ensure a matched melting temperature (Tm) [15]. |
Advanced Techniques for Primer-Dimer Suppression
Blocker Strands (Clamps): Short oligonucleotide strands can be added to the PCR mixture. They bind specifically to the primer-binding region of the template, blocking the primer from mishybridizing to off-target sites. This method suppresses errors through a combination of energetic destabilization and the creation of a kinetic barrier to mishybridization [17].
Self-Avoiding Molecular Recognition Systems (SAMRS): SAMRS involve primers with modified nucleobases (e.g., a, g, c, t). These SAMRS components pair normally with standard DNA (A:T, G:C) but do not pair with other SAMRS bases. This design significantly reduces primer-primer interactions, thereby preventing dimer formation and improving specificity in applications like SNP detection [6].
Experimental Protocols
Protocol 1: Optimizing Annealing Temperature Using a Gradient PCR
Purpose: To empirically determine the optimal annealing temperature (Ta) for a primer set to maximize specificity and yield while minimizing primer-dimer formation [15] [13].
- Prepare Master Mix: Create a standard PCR master mix containing template DNA, forward and reverse primers, dNTPs, reaction buffer, and a hot-start DNA polymerase.
- Aliquot: Dispense equal volumes of the master mix into multiple PCR tubes or a multi-well plate.
- Set Gradient: Program your thermal cycler's annealing step to use a temperature gradient that spans a range (e.g., 55°C to 70°C). The cycler will assign a different Ta to each column or row of tubes.
- Run PCR: Execute the PCR cycling program.
- Analyze Results: Visualize the PCR products using agarose gel electrophoresis. The optimal Ta is the highest temperature that produces a strong, specific band with the least or no primer-dimer [15].
Protocol 2: Evaluating Primer-Dimer Formation with a No-Template Control (NTC)
Purpose: To diagnose reagent contamination or confirm primer-dimer artifacts [2] [18].
- Setup: Prepare a standard PCR reaction mixture identical to your test samples, but replace the template DNA with nuclease-free water.
- Run in Parallel: Place the NTC tube in the thermal cycler alongside your experimental samples.
- Amplification: Run the full PCR protocol.
- Interpretation: Analyze the NTC result by gel electrophoresis. A clean NTC with no bands indicates no contamination. The presence of a band, particularly a smeary one below 100 bp, confirms primer-dimer formation and indicates that your reaction conditions require optimization [2].
Research Reagent Solutions
Table: Essential Reagents for Optimizing PCR and Reducing Primer-Dimer
| Reagent | Function & Rationale |
|---|---|
| Hot-Start DNA Polymerase | Remains inactive until a high-temperature activation step (e.g., 95°C), preventing enzymatic activity during reaction setup and thereby drastically reducing non-specific amplification and primer-dimer formation [2] [13]. |
| Magnesium Chloride (MgCl2) / Magnesium Sulfate (MgSO4) | An essential cofactor for DNA polymerase activity. Its concentration must be optimized, as low levels reduce enzyme activity, and high levels promote non-specific binding and primer-dimer formation [15] [13] [16]. |
| DMSO (Dimethyl Sulfoxide) | A PCR additive that helps denature DNA templates with strong secondary structures or high GC content, improving amplification efficiency and specificity. Typically used at 2-10% [15] [13]. |
| Betaine | An additive that homogenizes the base-pairing stability across DNA sequences, which is particularly useful for amplifying GC-rich templates. Often used at 1-2 M concentration [15]. |
| Uracil-DNA Glycosylase (UNG) | An enzyme added to PCR master mixes to prevent "carry-over" contamination from previous PCR products. It degrades uracil-containing DNA, which can be incorporated in place of thymine in previous reactions, thereby reducing false positives [18]. |
| Blocker Strands / Clamps | Short, modified oligonucleotides that bind to specific off-target sequences, blocking primers from mishybridizing. They suppress errors through both energetic and kinetic mechanisms [17]. |
Workflow and Relationship Diagrams
PCR Troubleshooting Workflow
Blocker Strand Mechanism
Definitions and Formation Mechanisms
Self-Dimerization occurs when two identical primer molecules anneal to each other due to regions of complementarity within a single primer sequence [2]. This intermolecular interaction can form homo-dimers [19].
Cross-Primer Dimerization involves two different primers (typically forward and reverse) annealing to each other due to complementary regions between them [2] [7]. This intermolecular interaction forms hetero-dimers [19].
Formation Process: Both mechanisms exploit complementarity between primer sequences. When primers contain regions that can base-pair with each other, they may bind together instead of to the template DNA, creating free 3' ends that DNA polymerase can extend [2]. This nonspecific amplification consumes reaction components and reduces target amplification efficiency [11].
Table 1: Key Characteristics of Primer Dimer Types
| Characteristic | Self-Dimerization | Cross-Primer Dimerization |
|---|---|---|
| Primers Involved | Two identical primers | Forward and reverse primers |
| Molecular Interaction | Intra-primer homology [7] | Inter-primer homology [7] |
| Dimer Type | Homo-dimer [19] | Hetero-dimer [19] |
| Common Cause | Regions of self-complementarity within a single primer [2] | Complementary sequences between different primers [2] |
| 3' End Complementarity | Fewer than 4 complementary bases recommended, especially at 3' end [19] | Fewer than 4 complementary bases recommended, especially at 3' end [19] |
Prevention Strategies and Primer Design Guidelines
Optimal Primer Design Parameters
Effective primer design represents the foremost strategy for preventing dimer formation [7]. Adhere to these critical parameters during design:
- Length: 18-30 nucleotides, with 18-24 being optimal [7] [20]
- GC Content: 40-60% [7] [20]
- Melting Temperature (Tm): 52-65°C, with both primers within 5°C of each other [7] [20]
- 3' End Stability: Include a G or C at the 3' end (GC clamp) but avoid more than 3 G/C bases consecutively [7] [20]
- Complementarity Check: Ensure fewer than 4 complementary bases, especially at the 3' ends [19]
Computational Design Tools
Leverage bioinformatics tools to automate compliance with these parameters [7]. Recommended platforms include:
- NCBI Primer-BLAST: Verifies target specificity and analyzes potential dimer formation [20]
- Primer3: Open-source tool for basic primer design [20]
- Eurofins Genomics Tools: Commercial solution with comprehensive dimer analysis [7]
Table 2: Quantitative Primer Design Guidelines for Minimizing Dimer Formation
| Design Parameter | Optimal Range | Rationale | Consequences of Deviation |
|---|---|---|---|
| Primer Length | 18-24 nucleotides [7] | Balances specificity and hybridization efficiency | Short primers: reduced specificity; Long primers: slower hybridization [7] |
| GC Content | 40-60% [7] [20] | Optimal hydrogen bonding stability | Low GC: weak binding; High GC: nonspecific binding & primer-dimers [7] |
| Melting Temperature (Tm) | 52-65°C [20], preferably 54°C or higher [7] | Ensures specific annealing | Low Tm: nonspecific binding; High Tm: secondary annealing [7] |
| 3' End Complementarity | <4 complementary bases [19] | Prevents polymerase extension of dimers | Primer-dimer formation and amplification [19] |
| Tm Difference Between Primers | ≤5°C [20] | Synchronized binding of both primers | Reduced amplification efficiency [20] |
Experimental Optimization Techniques
Reaction Component Adjustments
- Primer Concentration: Optimize between 0.1-1 μM; high concentrations promote dimer formation [13]
- Hot-Start DNA Polymerase: Utilizes polymerases inactive at room temperature, preventing pre-PCR extension and dimer formation during setup [2] [13]
- Magnesium Concentration: Optimize Mg²⁺ levels (typically 1.5-4.0 mM); excess Mg²⁺ promotes nonspecific amplification [13] [20]
Thermal Cycling Parameters
- Annealing Temperature: Increase temperature stepwise (1-2°C increments) to enhance specificity [2] [13]
- Denaturation Time: Increase denaturation times to disrupt primer interactions [2]
- Setup Temperature: Keep reactions on ice during preparation to minimize enzyme activity before thermal cycling [13]
Detection and Analysis Methods
Gel Electrophoresis Identification
Primer dimers exhibit characteristic features in agarose gel electrophoresis [2]:
- Short Length: Typically below 100 bp [2]
- Smeary Appearance: Fuzzy bands rather than well-defined bands [2]
- Mobility: Run gels longer to ensure dimers migrate past desired PCR products [2]
Control Reactions
Implement a No-Template Control (NTC) to identify primer-derived artifacts. Since primer dimers form without template DNA, they will appear as the primary amplification product in NTC reactions [2].
Research Reagent Solutions
Table 3: Essential Research Reagents for Primer Dimer Troubleshooting
| Reagent/Category | Function/Application | Usage Notes |
|---|---|---|
| Hot-Start DNA Polymerase | Remains inactive until high-temperature activation; minimizes pre-PCR extensions [2] [13] | Essential for multiplex PCR and low-template reactions |
| dNTPs | Deoxynucleotides (dATP, dCTP, dTTP, dGTP) for DNA synthesis [20] | Use balanced equimolar concentrations (typically 200μM each) |
| Magnesium Salts (MgCl₂/MgSO₄) | Cofactor for DNA polymerase; critical for reaction efficiency [13] [20] | Optimize concentration (0.5-5.0 mM); excess promotes nonspecificity |
| PCR Additives | DMSO, formamide, BSA, or betaine to improve specificity and reduce secondary structures [13] [20] | Use lowest effective concentration; adjust annealing temperature accordingly |
| Buffer Systems | Provides optimal pH and salt conditions for polymerase activity [20] | May contain Mg²⁺; check composition when calculating magnesium additions |
Advanced Troubleshooting Protocols
Systematic Optimization Workflow
Comprehensive Mitigation Strategy
When standard optimization proves insufficient, implement this hierarchical approach:
Redesign Primers: When complementarity cannot be resolved through optimization, redesign primers using computational tools to eliminate self-complementary regions and 3' overlaps [7] [19]
Advanced Polymerase Systems: Switch to specialized polymerase formulations with enhanced fidelity and specificity [13]
Touchdown PCR: Employ progressive annealing temperature reduction through cycles to enhance specificity in early amplification stages [13]
Additive Optimization: Systematically test PCR enhancers including DMSO (1-10%), formamide (1.25-10%), or betaine (0.5-2.5 M) to disrupt secondary structures [13] [20]
Successful PCR experimentation requires recognizing that primer dimer formation represents a common challenge rather than a procedural failure [2]. Through methodical implementation of these design principles, optimization strategies, and detection methods, researchers can effectively distinguish between self-dimerization and cross-primer dimerization mechanisms and implement targeted solutions to minimize their impact on experimental results.
Troubleshooting Guides
FAQ: How do primer dimers lead to reagent depletion and experimental delays?
Question: What are the direct economic and operational consequences of primer dimer formation in my PCR experiments?
Answer: Primer dimer formation directly impacts your research through two main channels:
- Reagent Depletion: Primer dimers are unintended amplification products that compete with your target DNA for essential, costly reagents. This nonspecific consumption drains your DNA polymerase, dNTPs, and primers, reducing the efficiency of your desired reaction and leading to failed experiments and the need for frequent reagent reorders [2] [6].
- Experimental Delays: The presence of primer dimers often results in failed or ambiguous experiments. This necessitates repetition of PCR runs, optimization procedures, and additional validation steps like gel electrophoresis. These activities consume valuable research time, delay project timelines, and reduce overall laboratory productivity [11].
This guide outlines a systematic approach to reduce primer dimer formation, saving both reagents and time.
Step 1: Interrogate Primer Design and Quality The most effective way to prevent primer dimers is to design and handle primers correctly.
- Action: Use primer design software to ensure primers have low self-complementarity and minimal 3'-end complementarity to each other [2] [20].
- Action: Check primer sequences for secondary structures like hairpin loops and avoid long runs of a single nucleotide [20].
- Action: Ensure primers are stored correctly in aliquots to prevent degradation and maintain quality [13].
Step 2: Optimize Reaction Components Fine-tuning your reaction mix can drastically reduce nonspecific amplification.
- Action: Lower primer concentrations to reduce the chance of primers annealing to each other. A typical working range is 0.1–1 μM [2] [13].
- Action: Use a hot-start DNA polymerase. These enzymes remain inactive until a high-temperature step, preventing nonspecific amplification during reaction setup [2] [21] [13].
- Action: Optimize Mg²⁺ concentration, as excess Mg²⁺ can promote non-specific binding and primer dimer formation [21] [13].
Step 3: Refine Thermal Cycling Conditions Adjusting your PCR protocol can enhance specificity.
- Action: Increase the annealing temperature incrementally. A higher temperature prevents primers from binding nonspecifically [2] [21].
- Action: Use a gradient PCR instrument to empirically determine the ideal annealing temperature for your primer set [21] [13].
- Action: Increase denaturation times to ensure DNA templates are fully separated [2].
The following workflow provides a visual summary of this systematic troubleshooting process:
Advanced Techniques for Stubborn Cases
For persistent primer dimer problems, especially in sensitive or multiplexed applications, consider these advanced strategies:
- Touchdown PCR: This technique starts with an annealing temperature above the primer's calculated Tm and gradually decreases it in subsequent cycles. This favors the amplification of the specific target when primers are most selective, effectively suppressing primer dimer formation [13] [6].
- Self-Avoiding Molecular Recognition Systems (SAMRS): SAMRS involves incorporating specially modified nucleobases into your primers. These SAMRS components pair normally with natural DNA but do not pair with each other. This technology can virtually eliminate primer-primer interactions, preventing dimer formation and is particularly valuable for highly multiplexed PCR and sensitive SNP detection [6].
The Scientist's Toolkit: Research Reagent Solutions
The table below lists key reagents and their optimized use for preventing primer dimers.
| Reagent/Item | Function & Optimization Guide for Reducing Primer Dimers |
|---|---|
| Primers | Function: Bind specifically to the target DNA sequence. Optimization: Design with biosoftware to avoid 3' complementarity. Use a concentration of 0.1-1 µM. High-quality, purified primers are essential [13] [20]. |
| Hot-Start DNA Polymerase | Function: Enzyme inactive at room temperature, preventing nonspecific amplification during setup. Optimization: Essential for minimizing pre-PCR primer-dimer formation. Choose antibodies or chemically modified versions for robust hot-start performance [2] [21] [13]. |
| Magnesium (Mg²⁺) | Function: Cofactor for DNA polymerase activity. Optimization: Concentration is critical. Excess Mg²⁺ promotes non-specific binding. Optimize in 0.2-1 mM increments from a typical starting point of 1.5 mM [21] [13]. |
| dNTPs | Function: Building blocks for new DNA strands. Optimization: Use balanced concentrations (200 µM of each dNTP is standard). Unbalanced dNTPs can increase error rate but are less directly linked to primer dimers [21] [13]. |
| Template DNA | Function: The target DNA to be amplified. Optimization: Use high-quality, pure template. Inhibitors or degraded DNA can reduce amplification efficiency, making primer dimers more apparent. Ensure correct quantity [13]. |
| PCR Additives | Function: Assist in amplifying difficult templates (e.g., GC-rich). Optimization: DMSO, Betaine, or BSA can help reduce secondary structures that might otherwise favor primer-dimer formation. Use at recommended concentrations [13] [20]. |
Experimental Protocols
Protocol 1: Standard PCR Setup with Hot-Start Polymerase
Objective: To set up a standard 50 µL PCR reaction designed to minimize primer dimer formation through the use of a hot-start polymerase and optimized cycling conditions.
Materials:
- Sterile, nuclease-free water
- 10X PCR Buffer (often supplied with Mg²⁺)
- dNTP Mix (10 mM total)
- Forward and Reverse Primers (20 µM stock each)
- Template DNA
- Hot-Start DNA Polymerase
- Thermal Cycler
Method:
- Prepare Master Mix: Thaw all reagents on ice. For multiple reactions, prepare a Master Mix in a sterile 1.5 mL tube to minimize pipetting errors and ensure consistency. Combine the following components in the order listed:
- Nuclease-free water (Q.S. to 50 µL)
- 10X PCR Buffer: 5 µL
- dNTP Mix (10 mM): 1 µL
- Forward Primer (20 µM): 1 µL
- Reverse Primer (20 µM): 1 µL
- Template DNA: 0.5-2 µL (e.g., 1-1000 ng)
- Hot-Start DNA Polymerase: 0.5-1.25 µL (follow manufacturer's instructions)
- Mix Thoroughly: Gently mix the reaction by pipetting up and down 20 times. Do not vortex.
- Run PCR: Place tubes in a pre-heated thermal cycler and start the following program:
- Initial Denaturation: 94–98°C for 2–5 minutes (activates hot-start polymerase).
- Amplification (25–35 cycles):
- Denature: 94–98°C for 20–30 seconds.
- Anneal: [Calculate Tm and test a gradient] 55–65°C for 20–30 seconds.
- Extend: 72°C for 1 minute per kb of amplicon.
- Final Extension: 72°C for 5–10 minutes.
- Hold: 4°C ∞.
- Analysis: Analyze 5-10 µL of the PCR product by agarose gel electrophoresis.
Notes: Always include a negative control (no template DNA) to detect contamination or primer dimer formation [2] [20].
Protocol 2: Optimization of Annealing Temperature Using a Gradient PCR
Objective: To empirically determine the optimal annealing temperature for a primer set to maximize specific product yield and minimize primer dimers.
Materials:
- Prepared PCR Master Mix from Protocol 1
- Thermal Cycler with gradient functionality
Method:
- Prepare Reactions: Aliquot your Master Mix into several PCR tubes.
- Set Gradient Program: Program your thermal cycler with a gradient across the annealing step. Set the range to span at least 5°C below and above the calculated Tm of your primers. For example, if the calculated Tm is 58°C, set a gradient from 53°C to 63°C.
- Run PCR: Start the cycling program.
- Analyze Results: Run the products on an agarose gel. Identify the temperature that produces the strongest band of your expected product size with the least amount of smearing or lower molecular weight bands (primer dimers). This temperature is your optimal annealing temperature for future experiments [21] [13].
Advanced Primer Design and Reaction Optimization Strategies
Frequently Asked Questions (FAQs)
Q1: What are the core principles for designing a high-quality PCR primer? The core principles involve optimizing four key characteristics: primer length, melting temperature ((T_m)), GC content, and 3'-end sequence. Adhering to these principles ensures specific binding to the target DNA and efficient amplification while minimizing side reactions like primer-dimer formation [7] [22].
Q2: How does primer design specifically influence primer-dimer formation? Primer-dimer formation is primarily caused by complementarity between primers, especially at their 3' ends [5]. If a primer has regions that are complementary to itself (self-dimer) or to the other primer in the pair (cross-dimer), they can anneal to each other instead of the template DNA. The DNA polymerase can then extend these bound primers, producing short, unintended products that compete with the target amplification and reduce PCR efficiency [11] [2].
Q3: What steps can I take if my PCR shows primer-dimer bands on a gel? If you observe primer-dimer (typically appearing as a fuzzy smear below 100 bp [2]), you can:
- Increase the annealing temperature to promote more specific primer binding [5] [2].
- Use a hot-start DNA polymerase, which is inactive until a high-temperature activation step, preventing spurious amplification during reaction setup [13] [2].
- Re-evaluate your primer design for complementarity [5].
- Lower the primer concentration in the reaction [13] [2].
- Run a no-template control (NTC) to confirm that the bands are indeed primer-dimers and not specific to your DNA sample [2].
Q4: Are there trusted tools to help me design primers and check for issues? Yes, several reliable tools are available:
- NCBI Primer-BLAST: This tool designs primers and checks their specificity against a database to ensure they only bind to your intended target [23].
- Primer3: A widely used open-source tool for selecting primers from a DNA sequence [5].
- NEB Tm Calculator: Helps evaluate the melting temperature and optimal annealing temperature for your primers, taking the specific polymerase and buffer into account [24].
Troubleshooting Guides
Problem 1: Non-Specific Amplification or Primer-Dimers
| Possible Cause | Recommended Solution | Experimental Protocol |
|---|---|---|
| Low annealing temperature | Increase the annealing temperature in 1-2°C increments [13] [5]. | Use a gradient PCR thermocycler to test a range of annealing temperatures (e.g., 50°C to 68°C) in a single run. The correct temperature will yield a single, strong band [5]. |
| Complementary primer sequences | Re-design primers to avoid self-complementarity or 3'-end complementarity between the forward and reverse primer [5] [25]. | Use primer analysis software to check the "self-complementarity" and "self 3'-complementarity" scores. A lower score is better [7]. |
| Excessive primer concentration | Titrate the primer concentration downwards [13] [2]. | Prepare a series of PCR reactions with primer concentrations ranging from 0.1 µM to 1 µM. Use the lowest concentration that provides a robust yield of the desired product [13] [9]. |
| Non-hot-start DNA polymerase | Switch to a hot-start enzyme [13] [2]. | Set up PCR reactions on ice and use a hot-start polymerase. Ensure the thermocycler has a pre-programmed initial denaturation step (e.g., 95°C for 2 minutes) to fully activate the enzyme before cycling begins. |
Problem 2: No Amplification or Low Yield
| Possible Cause | Recommended Solution | Experimental Protocol |
|---|---|---|
| Annealing temperature is too high | Lower the annealing temperature to facilitate primer binding [24]. | Perform a gradient PCR to find the optimal temperature, as described in Problem 1. |
| Poor primer specificity or quality | Verify primer specificity using BLAST and ensure high-quality, salt-free primers [13] [5]. | Order HPLC-purified primers. Use NCBI Primer-BLAST to confirm the primers are unique to your target sequence [23]. |
| Complex or GC-rich template | Use a polymerase and buffer system designed for GC-rich templates, and include additives [13] [24]. | Use a polymerase like Q5 High-Fidelity DNA Polymerase with its proprietary GC Enhancer. Alternatively, test additives like DMSO at a final concentration of 1-10% [24]. |
| Insufficient Mg²⁺ concentration | Optimize the Mg²⁺ concentration [13] [24]. | Prepare reactions with a gradient of MgCl₂ or MgSO₄ (e.g., from 1.0 mM to 4.0 mM in 0.5 mM increments) to identify the concentration that gives the highest yield [24]. |
Quantitative Data for Optimal Primer Design
The following table summarizes the target ranges for key primer parameters to minimize primer-dimer formation and ensure efficient amplification [7] [22] [25].
| Parameter | Optimal Range | Rationale & Technical Notes |
|---|---|---|
| Length | 18 - 30 nucleotides [22] [9] | Shorter primers bind more efficiently but longer primers offer greater specificity. 18-24 bases is often ideal for standard PCR [7]. |
| Melting Temperature ((T_m)) | 55°C - 65°C [25]; Pair (T_m) difference: ≤ 5°C [9] | (Tm) is the temperature at which 50% of the DNA duplex dissociates. Calculated using formulas like: (Tm = 4(G + C) + 2(A + T)) [7]. |
| GC Content | 40% - 60% [7] [22] [9] | GC bonds are stronger than AT bonds. Content within this range ensures stable binding without promoting secondary structures. |
| 3'-End Specificity (GC Clamp) | 1-2 G or C bases within the last 5 nucleotides [7] [22] | A "GC clamp" strengthens the binding of the critical 3' end, but more than 3 G/C bases can lead to non-specific binding [7]. |
Experimental Workflow for Primer Design and Validation
The diagram below outlines a systematic workflow for designing and validating primers, incorporating checks to reduce primer-dimer risk.
Research Reagent Solutions
The following table lists essential reagents and their specific functions in optimizing PCR and mitigating primer-dimer formation.
| Reagent / Material | Function & Role in Primer-Dimer Reduction |
|---|---|
| Hot-Start DNA Polymerase | Engineered to be inactive at room temperature. Prevents enzymatic extension of primerdimers formed during reaction setup. Activated only at high initial denaturation temperature [13] [2]. |
| High-Fidelity DNA Polymerase | Often possesses 3'→5' exonuclease (proofreading) activity, which can increase specificity and fidelity, reducing mispriming events that can lead to artifacts [9]. |
| GC Enhancer / Additives | Specialized buffers containing additives like DMSO, betaine, or glycerol. Help denature GC-rich templates and secondary structures, improving primer binding specificity and yield for difficult targets [13] [24]. |
| HPLC-Purified Primers | Purification method that removes short, truncated oligonucleotides. These truncated sequences can contribute to non-specific amplification and primer-dimer formation [5]. |
| MgCl₂ / MgSO₄ Solution | A crucial cofactor for DNA polymerase activity. Its concentration must be optimized, as excess Mg²⁺ can promote non-specific binding and primer-dimer formation [13] [24]. |
Leveraging Bioinformatics Tools for Assessing Self-Complementarity and Hairpin Formation
Frequently Asked Questions (FAQs)
1. What are self-complementarity and hairpin formation, and why are they problematic in PCR?
Self-complementarity occurs when regions within a single primer are complementary and can anneal to each other. Hairpin formation is a specific type of secondary structure where the primer folds back on itself, creating a loop and a double-stranded stem [2]. These structures are problematic because they prevent the primer from binding to its intended template DNA. This leads to reduced PCR efficiency, decreased yield of the desired product, and can complicate the interpretation of your results [2] [11].
2. Which key parameters should I check when analyzing my primer sequences?
When analyzing primers, you should evaluate several key thermodynamic parameters. The most critical ones are summarized in the table below.
| Parameter | Description | Optimal Range / Target |
|---|---|---|
| Self-Dimerization (ANY) [26] | Measure of a primer's tendency to anneal to itself. | As low as possible. |
| 3' Self-Complementarity [26] | Measure of tendency to form a primer-dimer at its 3' end. | 0 for maximum performance [26]. |
| Hairpin ΔG [26] | Change in free energy; negative value indicates a spontaneous (favorable) reaction. | Negative values are favorable [26]. |
| Melting Temperature (Tm) [27] | Temperature at which half of the DNA duplex dissociates. | Primer pair Tm difference (ΔTm) should be ≤ 5°C [26]. |
| GC Content [27] | Percentage of G and C bases in the primer. | Typically 40-60%. |
3. I have my primer sequences. What is the first tool I should use for analysis?
For an initial, comprehensive check, the Multiple Primer Analyzer from Thermo Fisher Scientific is an excellent starting point. This tool allows you to input multiple primer sequences simultaneously and instantly provides results for Tm, GC%, and a preliminary estimation of possible primer-dimers [27]. For more advanced specificity checking against genomic databases, NCBI's Primer-BLAST is the industry standard. It designs primers or checks pre-designed primers for specificity within a user-specified organism, helping to ensure your primers will bind only to the intended target [23].
4. My primer has a high "3' Self-Complementarity" score. What can I do?
A high 3' score is critical to address because the 3' end is where the polymerase extends. A mismatch or secondary structure here will severely interfere with synthesis [26]. You should strive for a score of 0. If your score is high, consider "moving" the primer sequence a few bases upstream or downstream on the template sequence and re-check the parameters [26]. If the core sequence cannot be altered, you can empirically add non-complementary G/C nucleotides to the 5' end to help stabilize the primer, but you must then re-check the new secondary structure [26].
5. Are there any specific experimental reagents that can help minimize issues from suboptimal primers?
Yes, several wet-lab reagents and techniques can mitigate problems. Hot-start DNA polymerases are highly recommended as they remain inactive until a high denaturation temperature is reached, minimizing primer-dimer formation during reaction setup [2]. You can also optimize your PCR buffer conditions, including Mg++ concentration, and use PCR enhancers to improve specificity [26]. Furthermore, lowering the primer concentration in the reaction can reduce opportunities for primers to anneal to each other rather than to the template [8] [11].
Troubleshooting Guides
Guide 1: A Step-by-Step Protocol for In silico Primer Analysis
Follow this detailed workflow to thoroughly analyze your primers before ordering them.
Procedure:
- Basic Quality Control Check: Use a tool like the Thermo Fisher Multiple Primer Analyzer [27]. Enter your forward and reverse primer sequences.
- Action: Verify that the Tm difference (ΔTm) between your primers is minimal (ideally ≤ 5°C). Check that GC% falls within an acceptable range (typically 40-60%).
- Output: Note the preliminary primer-dimer estimation provided by the tool.
- Check for Secondary Structures: Use tools like the Integrated DNA Technologies (IDT) OligoAnalyzer or mFold [26].
- Action: Input each primer sequence individually. For the OligoAnalyzer, click the 'Hairpin' tab to analyze the structure.
- Output: Retrieve the ΔG value from the analysis. A more negative ΔG indicates a more stable, and therefore more problematic, hairpin structure [26].
- Assess Primer Specificity: Use NCBI Primer-BLAST [23].
- Action: Input your primer sequences and select the appropriate organism under "Primer Pair Specificity Checking Parameters."
- Output: The tool will show all potential amplification products from the database, ensuring your primers are specific to your target gene and will not amplify unintended genomic regions [23].
- Holistic Parameter Visualization: Use a tool like PrimerChecker [26].
- Action: Input the thermodynamic parameters you have gathered (Tm, ΔG, ANY, 3' scores) for your primer pair.
- Output: PrimerChecker generates a radial plot that visually summarizes all parameters against optimal, good, and suboptimal ranges, allowing for rapid, holistic decision-making [26].
Guide 2: Troubleshooting Poor PCR Results
If your PCR results show a smeary band below 100 bp or low yield, follow this logical troubleshooting path.
Procedure:
- Run a No-Template Control (NTC): Include a control reaction in your PCR run that contains all reagents (water, buffer, primers, polymerase) except for the template DNA [2].
- Interpret the Results:
- If the NTC shows a smeary band around 100 bp, primer-dimer formation is confirmed. This means your primers are annealing to each other instead of the template.
- Execute Corrective Actions:
- Optimize Reaction Conditions:
- Redesign Primers: If optimization fails, the most effective long-term solution is to redesign your primers. Return to the Step-by-Step Protocol for In silico Primer Analysis and focus on achieving optimal parameters, particularly a 3' self-complementarity score of 0 [26].
Research Reagent Solutions
The following table lists key reagents and materials that are essential for implementing the protocols and troubleshooting steps outlined in this guide.
| Item | Function / Application |
|---|---|
| Hot-Start DNA Polymerase | Enzyme inactive at room temperature; minimizes primer-dimer formation during PCR setup [2] [11]. |
| Optimized PCR Buffers | Buffers with tailored MgCl₂ and additive concentrations; can enhance specificity and reduce mispriming [26]. |
| Nuclease-Free Water | Solvent for preparing reagent stocks; prevents enzymatic degradation of primers and templates. |
| Primer Design Software | Programs like Primer3 (integrated into Primer-BLAST) assist in selecting primer sequences with low self-complementarity [23] [2]. |
| Gel Electrophoresis System | For visualizing PCR products and identifying primer dimers as smeary bands below 100 bp [2]. |
Primer-dimer formation is a significant obstacle in polymerase chain reaction (PCR) experiments, consuming reagents and reducing the yield and specificity of target amplification. For researchers and drug development professionals, this challenge becomes even more critical in advanced applications like multiplex PCR and single-nucleotide polymorphism (SNP) detection. Chemical modifications to primer bases, such as Self-Avoiding Molecular Recognition Systems (SAMRS) and Locked Nucleic Acids (LNAs), offer powerful strategies to mitigate these issues by fundamentally altering primer-binding behavior. This guide provides troubleshooting advice and methodologies for effectively integrating these technologies into your experimental workflows to enhance PCR specificity.
Understanding the Technologies: SAMRS and LNAs
Self-Avoiding Molecular Recognition Systems (SAMRS)
SAMRS are synthetic nucleotide analogs engineered to bind complementarily to natural DNA but not to other SAMRS nucleotides. [28] This property, known as "self-avoidance," directly counters the primer-primer interactions that lead to dimer formation. [6] [29]
- Mechanism: SAMRS components (A, T, G, C) form stable pairs with their natural counterparts (A, T, G, C) via two hydrogen bonds, similar to a natural A:T pair. [6] [28] However, SAMRS:SAMRS pairs (e.g., A:T or G:C) are thermodynamically disfavored and form only weak, single hydrogen bonds, thus preventing stable binding between two SAMRS-containing primers. [6] [28] [29]
- Primary Application: The key application of SAMRS is in reducing or eliminating primer-dimer formation, which is especially valuable in highly multiplexed PCR assays where numerous primers are present simultaneously. [6] [29]
Locked Nucleic Acids (LNAs)
LNAs are modified RNA nucleotides characterized by a methylene bridge that "locks" the ribose ring in a specific conformation. [30] This lock enhances the base's binding affinity and thermal stability.
- Mechanism: The locked structure reduces conformational flexibility, pre-organizing the phosphate backbone for ideal binding to complementary sequences. [30] Each LNA base incorporated into a primer significantly increases its melting temperature (Tm). [31] [30]
- Primary Application: LNA modifications are primarily used to enhance the specificity and sensitivity of PCR assays. The increased Tm allows for the use of higher annealing temperatures, which discourages non-specific binding and primer-dimer formation. [31] [32] [30] They are particularly useful for amplifying difficult targets like GC-rich regions, short sequences (e.g., miRNA), and for discriminating SNPs. [30]
Table 1: Comparative Overview of SAMRS and LNA Technologies
| Feature | SAMRS | Locked Nucleic Acids (LNA) |
|---|---|---|
| Chemical Basis | Alternative nucleobases with altered hydrogen-bonding moieties [6] [28] | Modified RNA nucleotide with a methylene bridge [30] |
| Core Mechanism | Binds to natural DNA but not to other SAMRS bases [6] [29] | Increases duplex thermal stability (Tm) and binding affinity [31] [30] |
| Primary Benefit | Prevents primer-primer interactions and dimer formation [6] | Increases assay specificity and sensitivity; improves mismatch discrimination [31] [32] [30] |
| Typical Primer Design | Chimeric primers with SAMRS components strategically placed in the 3'-end [6] [29] | LNA monomers incorporated at specific positions, often near the 3' end or at mismatch sites [32] |
| Ideal for | Highly multiplexed PCR, SNP detection with low artifact formation [6] | SNP detection, short amplicons (miRNA), GC-rich targets, discriminating gene family members [30] |
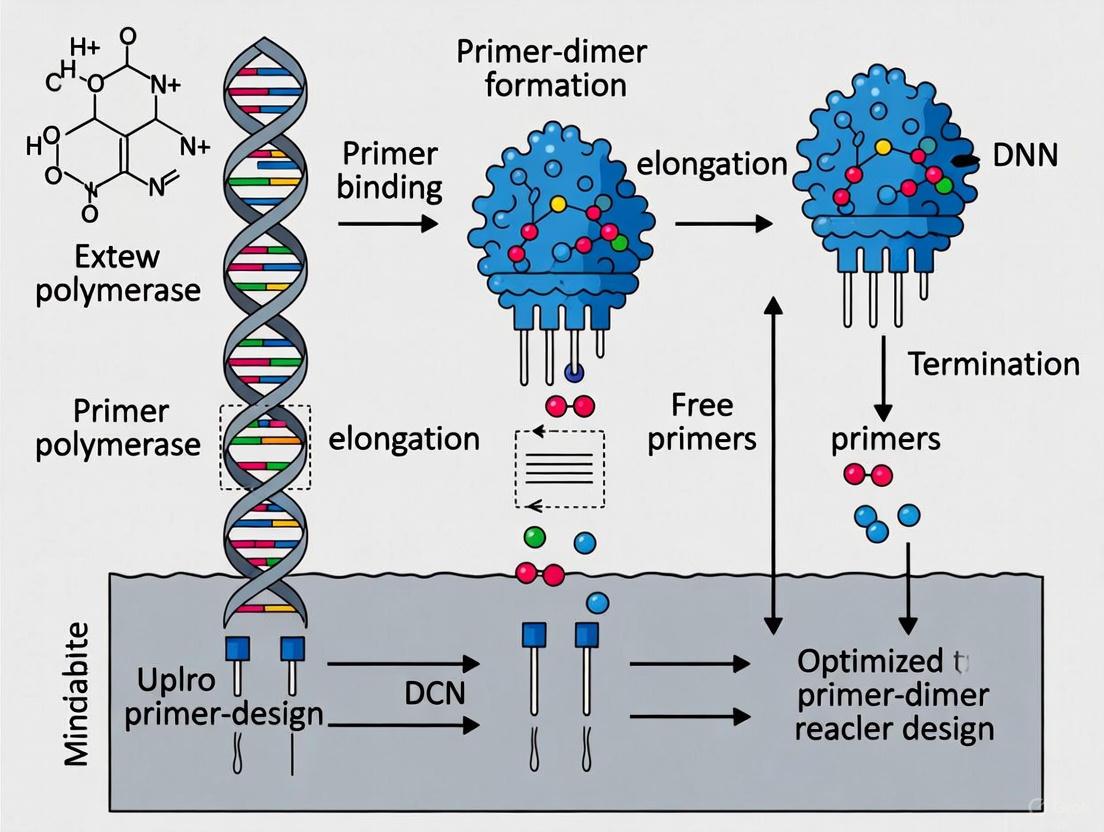
Diagram 1: Strategic pathways for enhancing PCR specificity using SAMRS and LNA.
Troubleshooting Guide: FAQs on SAMRS and LNA Implementation
FAQ 1: My multiplex PCR shows significant primer-dimer artifacts. How can SAMRS help?
- Problem: In multiplex PCR with many primer pairs, standard primers can interact, forming dimers that consume resources and obscure results. [6]
- Solution: Design chimeric primers with SAMRS components incorporated at their 3'-ends.
- Protocol:
- Primer Design: Synthesize chimeric primers in a {16+8+1} architecture: 16 standard nucleotides at the 5'-end, 8 SAMRS nucleotides (indicated by A, T, G, C*), and a single standard nucleotide at the 3'-end. [29] The 3'-terminal standard nucleotide is often retained to lower synthesis costs and maintain polymerase compatibility. [29]
- Experimental Setup: Use a hot-start DNA polymerase compatible with SAMRS, such as Taq DNA polymerase. [6] [29] Set up your multiplex PCR reaction as usual, replacing standard primers with the SAMRS-chimeric primers.
- Thermal Cycling: Use standard PCR cycling conditions appropriate for your target amplicons. The self-avoiding property of SAMRS is intrinsic and does not typically require cycling parameter adjustments. [6] [29]
- Expected Result: A significant reduction or elimination of primer-dimer bands on an agarose gel, with clean amplification of the desired multiplex targets. [29]
FAQ 2: How do I use LNAs to improve SNP discrimination in my assay?
- Problem: Standard primers fail to cleanly discriminate a single-nucleotide mismatch, leading to false-positive signals.
- Solution: Incorporate LNA bases into your primer at the position of the SNP to enhance the thermodynamic difference between a matched and mismatched binding event. [30]
- Protocol:
- Primer Design: Design primers where the LNA modification is placed at the SNP site or within the last few bases at the 3' end. [32] This placement maximizes the discriminatory power, as an LNA-induced increase in Tm is only fully realized with a perfectly matched base. [32] [30]
- Optimization: It is often necessary to design several primers with the LNA at different positions and test them empirically. [32] For example, test a forward primer with an LNA at the first base from the 3' end, another with an LNA three bases in, and a third with LNAs in both positions. [32]
- Thermal Cycling: Precisely optimize the annealing temperature. The increased Tm provided by the LNA allows you to use a higher, more stringent annealing temperature, which will further suppress amplification from the mismatched template. [30]
- Expected Result: Successful amplification only from the perfectly matched template, with little to no signal from the template containing the SNP. [30]
FAQ 3: My PCR has low yield and specificity despite using LNAs. What am I doing wrong?
- Problem: Poor results with LNA-modified primers are often due to suboptimal design.
- Troubleshooting Steps:
- Verify LNA Placement: Incorrect placement of LNA bases can lead to reduced specificity or even increased non-specific binding. [30] Avoid clustering too many LNA bases; typically, a few strategically placed modifications are more effective than a fully modified primer.
- Check Annealing Temperature: The incorporation of LNAs raises the primer's Tm. If the annealing temperature is not adjusted upward accordingly, you will lose the specificity benefit. Use a thermal gradient cycler to determine the optimal annealing temperature for your LNA primer. [30]
- Assess Primer Quality: Ensure oligonucleotides were synthesized with purification to remove truncated products, which can contribute to background noise. [13]
Essential Research Reagent Solutions
The successful implementation of SAMRS and LNA technologies relies on key reagents and design considerations.
Table 2: Key Reagents and Materials for Experimental Workflows
| Reagent / Material | Function & Importance | Considerations for Use |
|---|---|---|
| SAMRS Phosphoramidites | Building blocks for the chemical synthesis of SAMRS-containing oligonucleotides. [6] | Source from specialized manufacturers (e.g., Glen Research, ChemGenes). Requires standard phosphoramidite chemistry without special coupling conditions. [6] |
| LNA-Modified Oligonucleotides | Primers or probes with enhanced binding affinity and specificity. [31] [30] | Order from vendors that guarantee precise incorporation. LNA bases are often denoted in sequences by underlining, brackets, or a '+' prefix. [32] |
| Hot-Start DNA Polymerase | A polymerase inactive at room temperature, preventing non-specific priming and primer-dimer formation during reaction setup. [6] [2] [13] | Essential for maximizing the benefits of both SAMRS and LNA, as it suppresses artifacts before thermal cycling begins. |
| Compatible DNA Polymerase | An enzyme that efficiently incorporates nucleotides from SAMRS or LNA primers. | Taq DNA polymerase has been shown to work well with SAMRS components. [29] Verify polymerase compatibility with LNA primers for robust amplification. |
| HPLC Purification | A purification method for synthetic oligonucleotides to ensure high purity and correct sequence length. [6] | Critical for both SAMRS and LNA primers. Impure primers (e.g., containing truncated sequences) are a major source of non-specific amplification and failed experiments. [6] [13] |
Diagram 2: Molecular structures and binding relationships of SAMRS and LNA.
Core Principles: The Interplay of Key Components
How do primer concentration, Mg2+, and additives interact within a Master Mix to influence primer-dimer formation?
The core components of a PCR Master Mix do not function in isolation; their interactions determine the reaction's specificity. Primer-dimer formation occurs when primers anneal to each other instead of the target DNA template, largely due to complementary sequences at their 3' ends [11]. The balance of components either suppresses or promotes this:
- Primer Concentration: Excessive primer concentrations increase the probability of primers encountering and binding to each other, rather than to the template [33] [34]. Reducing primer concentration is a primary strategy to lower this probability [33].
- Mg2+ Concentration: Magnesium ions are essential cofactors for DNA polymerase activity. However, excessively high concentrations can stabilize the weak, non-specific bonds between primer molecules, facilitating primer-dimer formation and extension [13] [34].
- Chemical Additives: Additives like DMSO, betaine, and formamide can alter the stringency of primer annealing. They help by destabilizing secondary structures or reducing the effective melting temperature (Tm), thereby promoting specific primer-template binding over spurious primer-primer interactions [20] [13].
Optimizing a Master Mix involves systematically adjusting these components to create conditions where specific primer-template binding is overwhelmingly favored.
Troubleshooting Guide: Common Master Mix Issues and Solutions
| Observed Problem | Potential Cause | Recommended Solution |
|---|---|---|
| Primer-dimer formation | High primer concentration; Excess Mg2+; Low annealing temperature [11] [13] [33] | Optimize primer concentration (0.1–1 µM); Titrate Mg2+ downward; Increase annealing temperature [13] [33]. |
| No amplification or low yield | Insufficient Mg2+; Suboptimal primer concentration; Missing additives for complex templates [13] [34] | Titrate Mg2+ upward (0.5-5.0 mM); Verify primer concentration; Include 1-10% DMSO or 0.5-2.5 M betaine for GC-rich targets [20] [13]. |
| Non-specific amplification | Excess Mg2+; Low annealing temperature; High primer concentration [13] [35] | Reduce Mg2+ concentration; Increase annealing temperature; Use hot-start DNA polymerase [13] [34]. |
| Inconsistent results | Non-homogeneous Master Mix; Degraded reagents [13] | Mix all reagent stocks thoroughly before use; Prepare fresh working aliquots of critical components [13]. |
Optimizing Master Mix Components: Quantitative Guidelines and Protocols
Primer Concentration Optimization
Primer concentration is a critical variable. While standard protocols often use 0.5 µM, optimization between 0.1 µM and 1 µM is recommended [13] [35]. High concentrations promote primer-dimer formation, but insufficient concentration leads to poor amplification efficiency [34].
Protocol: Primer Titration Experiment
- Prepare a Master Mix containing all standard components except primers.
- Aliquot the Master Mix into separate tubes.
- Add forward and reverse primers to each tube to achieve final concentrations of 0.1, 0.2, 0.5, and 1.0 µM.
- Run the PCR using your standard cycling conditions.
- Analyze the results by gel electrophoresis. The optimal concentration yields the strongest specific band with the least primer-dimer.
Mg2+ Concentration Optimization
Mg2+ is a crucial cofactor, and its optimal concentration can vary with primer-template combination and the presence of chelators like EDTA [13]. Most protocols start with 1.5 mM, but a titration is often necessary.
Protocol: Mg2+ Titration Experiment
- Prepare a Master Mix without Mg2+.
- Aliquot the Master Mix into separate tubes.
- Add MgCl2 or MgSO4 (check polymerase preference [13]) to achieve a final concentration series (e.g., 0.5, 1.0, 1.5, 2.0, 3.0, 4.0 mM).
- Run the PCR and analyze the products. Select the lowest concentration that gives robust, specific amplification [20] [35].
Selecting and Using Additives
Additives are particularly useful for amplifying difficult templates, such as those with high GC content or complex secondary structures.
Reference Table: Common PCR Additives
| Additive | Common Final Concentration | Primary Function | Considerations |
|---|---|---|---|
| DMSO | 1 - 10% [20] | Disrupts DNA secondary structures, lowers Tm [13]. | High concentrations can inhibit polymerase; requires adjustment of annealing temperature [13]. |
| Betaine | 0.5 M - 2.5 M [20] | Equalizes nucleotide stability, helps denature GC-rich regions [13]. | Often used for high-GC templates. |
| Formamide | 1.25 - 10% [20] | Denaturant that increases stringency. | Can be inhibitory; use at lower concentrations. |
| BSA (Bovine Serum Albumin) | 10 - 100 μg/ml [20] | Binds to inhibitors in the reaction, stabilizing polymerase [34]. | Useful when template purity is suspect. |
Advanced Strategies: Hot-Start Polymerases and In Silico Design
Utilizing Hot-Start DNA Polymerases
Hot-start polymerases are inactive at room temperature, preventing enzymatic activity during reaction setup—a common time for primer-dimer formation. They are activated only after a high-temperature denaturation step [13] [34]. Incorporating a hot-start polymerase is one of the most effective ways to reduce non-specific amplification and primer-dimer artifacts [34].
Computational Primer Design and Prediction
Modern primer design leverages software to minimize inherent complementarity that leads to dimerization [20] [33]. Tools like Primer3 and NCBI Primer-Blast are standard for checking self-complementarity and off-target binding [20]. Advanced methods are now using machine learning, such as recurrent neural networks (RNNs), to predict PCR success or failure from primer and template sequences, potentially reducing reliance on extensive empirical optimization [36].
Frequently Asked Questions (FAQs)
Q1: What is the single most important adjustment to reduce primer-dimer? A: There is no single solution, but a combination approach is best. Start by ensuring good primer design with minimal self-complementarity. Then, empirically optimizing the primer concentration and using a hot-start polymerase often yield the most significant improvements [11] [33] [34].
Q2: Can I simply keep reducing primer concentration until the dimer disappears? A: No. Excessively low primer concentration will lead to inefficient amplification or no product [34]. A titration within the 0.1–1 µM range is necessary to find the optimal balance between yield and specificity [13].
Q3: How does Mg2+ specifically contribute to primer-dimer formation? A: Mg2+ stabilizes all double-stranded nucleic acid interactions, including the specific primer-template complex and the non-specific primer-primer complex. High Mg2+ concentrations can provide enough stability to transient primer-primer hybrids, allowing the DNA polymerase to extend them into a stable primer-dimer product [13] [34].
Q4: When should I consider using additives like DMSO or betaine? A: Additives are particularly helpful when amplifying GC-rich templates (>60% GC) or templates with pronounced secondary structures. If optimization of primer concentration, Mg2+, and annealing temperature fails, introducing a low concentration of an additive (e.g., 3-5% DMSO) can be highly effective [20] [13].
The Scientist's Toolkit: Essential Research Reagent Solutions
| Reagent / Material | Critical Function in Master Mix Optimization |
|---|---|
| Hot-Start DNA Polymerase | Prevents non-specific amplification and primer-dimer formation during reaction setup by remaining inactive until a high-temperature step [13] [34]. |
| Ultra-Pure dNTPs | Provides the essential nucleotides for DNA synthesis. Unbalanced or impure dNTPs can increase error rate and reduce amplification efficiency [13]. |
| MgCl2 or MgSO4 Stock | The source of Mg2+ ions. The type of salt (Cl vs. SO4) can affect polymerase performance and should be selected based on manufacturer recommendations [13]. |
| PCR-Grade Water | A nuclease-free, sterile water is essential to avoid degradation of primers and templates and to prevent introduction of PCR inhibitors. |
| Chemical Additives (DMSO, Betaine) | Used to modify reaction stringency and assist in denaturing complex DNA templates, thereby improving specificity and yield for challenging targets [20] [13]. |
| Gradient Thermal Cycler | Allows for the empirical testing of a range of annealing temperatures in a single run, drastically speeding up the optimization process [13]. |
FAQs and Troubleshooting Guides
What is the primary cause of primer-dimer formation, and how do optimized annealing temperatures help?
Primer-dimers are short, double-stranded DNA artifacts that form when PCR primers anneal to each other instead of to the target DNA template. This occurs primarily due to complementary sequences within or between the primers, especially at their 3' ends, and is facilitated by low stringency conditions during PCR setup and the initial cycling phases [7] [37] [11].
Optimizing the annealing temperature is a fundamental strategy to enhance specificity. Using an annealing temperature that is too low allows primers to bind to unintended, partially complementary sequences or to each other, leading to primer-dimer formation and nonspecific amplification. A higher, optimized annealing temperature promotes stringent binding, ensuring primers anneal only to their perfect complementary target sequence [7] [38].
How do I calculate and optimize the annealing temperature for my primer set to prevent primer-dimers?
The annealing temperature (Ta) is directly based on the melting temperature (Tm), which is the temperature at which 50% of the primer-DNA duplex dissociates [7]. The goal is to use a Ta that is high enough for specificity but not so high that the primer cannot bind.
Calculation and Optimization Guidelines:
- Melting Temperature (Tm): Aim for primer Tm values between 55°C and 75°C [39] [22]. Both primers in a set should have Tm values within 5°C of each other [22] [20].
- Initial Ta Estimate: A common starting point is to set the Ta at 2–5°C below the Tm of the primers [7].
- Empirical Optimization: The calculated Tm is a theoretical starting point. The optimal Ta should be determined experimentally using a gradient thermal cycler, testing a range of temperatures (e.g., 50°C to 70°C) to identify the temperature that yields the highest target product yield with the lowest primer-dimer formation [39].
- Universal Annealing: Some specialized DNA polymerases and buffer systems are designed to allow for a universal annealing temperature of 60°C, simplifying protocol setup and reducing optimization time without compromising specificity [39].
Table 1: Key Primer Design Parameters to Minimize Primer-Dimers
| Parameter | Recommended Range | Rationale |
|---|---|---|
| Primer Length | 18–30 nucleotides [7] [22] [20] | Balances specificity and efficient binding. |
| GC Content | 40–60% [7] [22] [20] | Prevents overly weak (low GC) or strong (high GC) binding that can lead to mismatches. |
| GC Clamp | Presence of G or C at the 3'-end [7] [22] | Strengthens binding at the critical point of polymerase extension; avoid >3 G/C in the last 5 bases [7]. |
| Melting Temp (Tm) | 55–75°C; primers within 5°C [39] [22] [20] | Ensures both primers bind with similar efficiency at the same Ta. |
| Self-Complementarity | Avoid repeats of >3 bases and inter-primer homology [7] [22] | Minimizes chances of hairpins and primer-self-dimer or cross-dimer formation. |
What is Hot-Start activation, and how does it reduce primer-dimer formation?
Hot-Start activation is a technique that inhibits DNA polymerase activity during the reaction setup and initial heating phases, which occur at lower, non-stringent temperatures. This prevents the polymerase from extending primers that are bound to non-specific targets or to each other, a primary cause of primer-dimer accumulation [40] [37].
Mechanism of Action: At room temperature, primers can transiently bind to off-target sequences or other primers via short complementary regions. Without Hot-Start activation, the polymerase can extend these misprimed complexes, synthesizing unwanted products that then compete for reagents in subsequent cycles. Hot-Start modifications keep the polymerase inactive until a high-temperature activation step (e.g., 95°C for initial denaturation) is reached, ensuring the reaction begins with a "hot start" under stringent conditions [40].
What are the different types of Hot-Start technologies, and how do I choose?
Hot-Start technologies employ various mechanisms to temporarily inhibit the DNA polymerase. The choice depends on the required stringency, activation time, and experimental constraints.
Table 2: Comparison of Common Hot-Start PCR Technologies
| Hot-Start Technology | Mechanism | Benefits | Considerations |
|---|---|---|---|
| Antibody-Based [40] | An antibody binds the polymerase's active site, blocking activity until the initial denaturation step inactivates the antibody. | Fast activation; full enzyme activity restored; similar performance to non-hot-start versions. | May contain animal-origin components. |
| Chemical Modification [40] | Polymerase is covalently modified with a chemical group that blocks activity. | High level of stringency at low temperatures. | Requires longer initial activation time (e.g., 5-10 minutes); may not be ideal for very long amplicons. |
| Affibody/Aptamer-Based [40] | A small protein (Affibody) or oligonucleotide (Aptamer) binds and blocks the polymerase. | Short activation time; often free of animal-origin components. | May be less stringent than antibody-based methods. |
| Primer-Based [37] | The primers themselves are chemically modified at the 3'-end (e.g., with a thermolabile group) to block extension. | High specificity as the modification is directly on the primer. | Requires custom synthesized primers; additional cost. |
Hot-Start PCR Activation Workflow
My PCR still shows primer-dimers after optimizing annealing temperature and using Hot-Start. What else can I troubleshoot?
Several other factors in the reaction setup and thermal cycling protocol can be adjusted to further suppress primer-dimer formation.
1. Reagent Quality and Concentration:
- Primer Concentration: Using excessively high primer concentrations increases the likelihood of primer-primer interactions. Titrate primer concentrations, typically between 0.1–0.5 µM each, to find the lowest concentration that still provides robust amplification [20].
- Template Quality and Quantity: Contaminants or an suboptimal amount of template DNA can promote nonspecific amplification. Use high-quality, pure DNA template. The optimal amount is usually 10 pg–100 ng of genomic DNA, depending on the template complexity and target abundance [38].
2. Thermal Cycler Protocol Adjustments:
- Touchdown PCR: This method starts with an annealing temperature higher than the calculated Tm and gradually decreases it over subsequent cycles. This ensures that the first, most critical cycles are of the highest stringency, favoring the amplification of the specific target over primer-dimers [38].
- Pre-incubation on Ice: Always prepare the PCR master mix on ice to minimize enzyme activity and primer interactions before the reaction begins [40].
- Protocol Type: If the primer Tm is high enough (e.g., >65°C), consider a two-step PCR protocol that combines the annealing and extension steps at 68–72°C, eliminating the low-temperature annealing phase entirely [38].
3. Reaction Additives: Certain additives can help improve specificity by altering DNA duplex stability or polymerase processivity. A common example is DMSO (1–10%), which is particularly helpful for amplifying GC-rich templates that are prone to forming secondary structures [20] [38]. Other additives include formamide, betaine, and BSA [20].
Troubleshooting Paths for Persistent Primer-Dimers
Research Reagent Solutions
The following table lists key reagents and their specific functions in optimizing PCR to reduce primer-dimer formation.
Table 3: Essential Reagents for PCR Optimization
| Reagent / Tool | Specific Function in Reducing Primer-Dimers |
|---|---|
| Hot-Start DNA Polymerase | Suppresses non-specific amplification during reaction setup by remaining inactive until a high-temperature step [40]. |
| Gradient Thermal Cycler | Empirically determines the optimal annealing temperature by running simultaneous reactions across a temperature range [39]. |
| Primer Design Software (e.g., NCBI Primer-BLAST) | Automates the design of specific primers with appropriate length, Tm, and GC content while checking for self-complementarity [20]. |
| DMSO (Dimethyl Sulfoxide) | Additive that disrupts secondary structures and can improve hybridization specificity, especially for GC-rich templates [20] [38]. |
| Magnesium Chloride (MgCl₂) | Critical cofactor for polymerase activity; its concentration can be optimized (e.g., 1.5–2.5 mM) to enhance fidelity and reduce mispriming [20] [38]. |
Practical Laboratory Troubleshooting: Systematic Problem-Solving for Primer-Dimer Issues
Troubleshooting Guides
Why is there amplification in my No-Template Control (NTC)?
Problem: Your No-Template Control (NTC) shows amplification, either as a band on a gel or a Ct value in qPCR, indicating that your reaction is amplifying something other than your intended target.
Investigation: The first step is to determine the cause. The investigation and solutions depend on whether the issue is contamination or primer-dimer formation. The flowchart below outlines the diagnostic process.
Solutions for Contamination
If you have diagnosed your problem as contamination, follow this systematic protocol to clean up your lab space and processes [41] [42].
- Physical Separation of Workspaces: Establish and strictly enforce separate "pre-PCR" and "post-PCR" areas [41] [42]. The pre-PCR area, ideally a dedicated hood with a UV light, is used only for preparing master mixes and must never contain template DNA or amplified PCR products. The post-PCR area is for all work involving templates and analyzed PCR products.
- Use Dedicated Equipment and Supplies: Use a dedicated set of pipettes, exclusively for the pre-PCR area. Always use aerosol-resistant (filter) pipette tips to prevent contamination of the pipette barrel [42].
- Reagent Management: Aliquot all reagents (polymerase, dNTPs, primers, water) into single-use volumes upon arrival to minimize the risk of contaminating entire stocks [42].
- Decontaminate Your Workspace: Before starting a new experiment, wipe down all surfaces, pipettes, and equipment with a 10% bleach solution or a commercial DNA decontaminant (e.g., DNA-Away). If available, expose your PCR hood to UV light for 15-30 minutes to degrade any contaminating DNA [42].
- Enzymatic Control: For recurring carryover contamination from previous PCR products, incorporate Uracil-N-Glycosylase (UNG) or Uracil-DNA Glycosylase (UDG) into your master mix. This enzyme degrades uracil-containing DNA (from previous dUTP-containing PCRs) before the thermal cycling starts, preventing its amplification [41].
Solutions for Primer-Dimer Formation
If you have diagnosed your problem as primer-dimer formation, the following optimization strategies can help.
- Optimize Primer Design: This is the most effective long-term solution. Redesign primers using trusted software to ensure they have low self-complementarity and cross-complementarity, especially at the 3' ends. Avoid more than 3 G/C bases in the last 5 nucleotides at the 3' end [7] [5].
- Increase Annealing Temperature: A higher annealing temperature promotes specific primer binding and reduces the chance of primers loosely annealing to each other. Optimize by testing a temperature gradient from 3–5°C below to 3–5°C above the calculated primer Tm [2] [13] [5].
- Use Hot-Start DNA Polymerase: Hot-start polymerases remain inactive until the initial high-temperature denaturation step. This prevents enzymatic activity during reaction setup at room temperature, when primer-dimer formation is most likely to initiate [2] [13] [43].
- Optimize Primer Concentration: High primer concentration increases the likelihood of primers interacting. Titrate primer concentrations downwards (e.g., from 1 µM to 0.1 µM) to find the lowest concentration that still provides robust amplification of your target [41] [13] [5].
- Adjust Thermal Cycler Protocol: Ensure the reaction is kept on ice during setup and transferred immediately to a pre-heated thermal cycler. You can also use a "hot start" protocol where the cycler pre-heats to 95°C before the block is accessed [5].
Why are my gel bands smeared or poorly resolved?
Problem: Instead of sharp, distinct bands, your gel shows smeared, fuzzy, or poorly separated bands, making interpretation difficult.
Investigation and Solutions:
- Gel Preparation:
- Cause: Gels thicker than 5mm can cause band diffusion. Poorly formed or damaged wells can lead to sample leakage and smearing [44].
- Solution: Cast horizontal agarose gels to a thickness of 3–4 mm. Use clean combs, avoid pushing them to the very bottom of the gel tray, and remove them carefully after the gel has solidified completely [44].
- Sample Preparation:
- Cause: Overloading the well with too much DNA, using degraded DNA, or having a sample in a high-salt buffer can all cause smearing [44] [42].
- Solution:
- Quantity: Do not exceed 0.1–0.2 µg of DNA per millimeter of well width [44].
- Degradation: Use molecular biology grade reagents and nuclease-free labware. Run an aliquot of your template DNA on a gel to check for integrity before PCR [44] [13].
- Purity: If the sample is in a high-salt buffer, dilute, precipitate, or re-purify the DNA and resuspend in nuclease-free water [44].
- Gel Run Conditions:
- Cause: Applying very low or very high voltage, or running the gel for an insufficient or excessively long time, leads to suboptimal resolution [44].
- Solution: Apply the voltage recommended for your gel percentage and nucleic acid size. Ensure the run time is long enough to resolve bands but not so long that excessive heat causes diffusion [44].
Frequently Asked Questions (FAQs)
Q1: What does a single, sharp band in my sample lane mean? A: This is typically the ideal result in a PCR experiment. It indicates that your primers have successfully and specifically amplified a single target DNA sequence of the expected size. You should confirm the size by comparing it to a DNA ladder [42].
Q2: My negative control is clear, but my sample lanes show a bright, fast-migrating band at the bottom of the gel. What is it? A: This is highly indicative of primer-dimer. It will appear as a fuzzy band or smear below 100 bp. Because it doesn't require a template, it can form in any reaction (including your NTC, but sometimes it's only visible in samples if the conditions are just at the threshold). You should still troubleshoot to minimize it, as it can compete with your target amplification and reduce yield [2].
Q3: How can I definitively confirm if the band in my NTC is a primer-dimer or contamination? A: For SYBR Green qPCR users, run a dissociation (melt) curve analysis after amplification. Primer-dimers will produce a distinct peak at a lower melting temperature (Tm) than your specific PCR product [41]. For conventional PCR users, the most straightforward method is to compare the band size to your target product on a gel. A much smaller size points to primer-dimer. If possible, you can also sequence the band from the NTC for a definitive answer.
Q4: What is the single most important practice to prevent contamination? A: The most critical practice is physical separation of pre-PCR and post-PCR activities and using dedicated equipment and filter tips for each area. This prevents amplified DNA from previous experiments from being introduced into new reactions [42].
Research Reagent Solutions
The following reagents are essential for implementing the troubleshooting strategies discussed above.
| Reagent/Material | Function in Troubleshooting | Key Consideration |
|---|---|---|
| Hot-Start DNA Polymerase | Reduces primer-dimer formation by remaining inactive until the initial denaturation step [2] [13]. | Choose enzymes with proven hot-start capability (e.g., antibody-mediated or chemical modification). |
| Uracil-N-Glycosylase (UNG) | Prevents carryover contamination by degrading PCR products from previous reactions that contain dUTP [41]. | Requires the use of dUTP in place of dTTP in all PCR master mixes. |
| HPLC-Purified Primers | Ensures high-quality primers with minimal truncated sequences, reducing non-specific binding and dimer formation [5]. | Specify HPLC purification when ordering oligonucleotides. |
| Nuclease-Free Water | Serves as a pure, uncontaminated base for preparing reagents and master mixes [42]. | Always aliquot from a fresh, certified stock bottle. |
| Aerosol-Resistant Filter Tips | Prevents pipette barrel contamination by aerosols, a major source of cross-contamination [42]. | Non-negotiable for PCR setup; use in both pre- and post-PCR areas. |
| DNA Decontamination Solution | Degrades contaminating DNA on lab surfaces and equipment [42]. | A 10% bleach solution or commercial products like DNA-Away are effective. |
Experimental Protocols
Protocol: Optimizing Primer Concentration to Minimize Dimer Formation
This protocol systematically tests different primer combinations to find the concentration that provides specific amplification with minimal primer-dimer [41].
- Prepare a Master Mix without primers, calculated for 9 reactions. Include all other components: buffer, dNTPs, hot-start polymerase, nuclease-free water, and template DNA.
- Aliquot the master mix into 9 PCR tubes.
- Add Primers according to the following matrix. The notation Forward/Reverse indicates the concentration in nM (e.g., 100/200 = 100 nM Forward Primer, 200 nM Reverse Primer).
- Run PCR using a previously established or optimized thermal cycling protocol.
- Analyze Results by running the products on a 2-3% agarose gel. Identify the well with the strongest target band and the faintest or absent primer-dimer band.
Table: Primer Concentration Optimization Matrix
| Reverse Primer (nM) | 100 nM Forward | 200 nM Forward | 400 nM Forward |
|---|---|---|---|
| 100 nM | 100/100 | 200/100 | 400/100 |
| 200 nM | 100/200 | 200/200 | 400/200 |
| 400 nM | 100/400 | 200/400 | 400/400 |
Protocol: Performing a Gradient PCR for Annealing Temperature Optimization
This protocol helps determine the ideal annealing temperature (Ta) for your specific primer set, which is crucial for specificity [13] [5].
- Prepare a Master Mix containing all reaction components, including primers (use a mid-range concentration from the optimization above, e.g., 200 nM each) and template.
- Aliquot the same volume of master mix into each tube of a gradient thermal cycler.
- Set the Gradient: Program your thermal cycler's annealing step to a temperature gradient that spans a realistic range (e.g., from 50°C to 68°C). The cycler will automatically assign a different Ta to each tube.
- Run the PCR.
- Analyze Results by gel electrophoresis. Load all reactions side-by-side. You are looking for the highest temperature that yields a strong, specific band with no non-specific products or primer-dimer. This temperature represents the optimal balance between yield and specificity.
FAQ: Troubleshooting Primer-Dimer Formation
1. What is a primer-dimer and how does it affect my PCR? A primer-dimer is a small, unintended DNA fragment that forms when PCR primers anneal to each other instead of to the target DNA template [2]. This occurs due to complementary regions between primers (cross-dimer) or within a single primer (self-dimer) [7] [10]. Primer-dimers consume reaction reagents—primers, nucleotides, and polymerase activity—which reduces the efficiency and yield of your target amplification [5] [10]. In quantitative PCR (qPCR), this can lead to false positives, false negatives, or inaccurate quantification [45] [10].
2. Why is annealing temperature so critical for preventing primer-dimer? Primer-dimers form most readily at low annealing temperatures, where the weak complementary binding between primers is stable [2] [5]. A higher annealing temperature promotes stringent binding, ensuring primers only anneal to their perfectly matched target sequences [2] [11]. Using an annealing temperature that is too low allows primers to bind to non-target sequences and to each other, dramatically increasing the risk of primer-dimer formation and other non-specific products [5].
3. How does a temperature gradient help optimize my PCR assay? A temperature gradient experiment allows you to test a range of annealing temperatures simultaneously on the same thermal cycler [5]. This is the most efficient way to empirically determine the optimal annealing temperature for your specific primer-template combination. By comparing the results across temperatures, you can identify the "sweet spot"—the highest temperature that yields robust, specific amplification of your target with little to no primer-dimer [5].
4. My primers were designed with software. Do I still need to optimize the annealing temperature? Yes. In silico calculations of melting temperature (Tm) provide a theoretical starting point, but the optimal annealing temperature (Ta) can vary in practice due to factors like specific buffer composition, enzyme formulation, and template quality [7] [20]. Empirical testing via a temperature gradient is the only way to confirm the most specific and efficient conditions for your actual reaction setup [45].
5. What other factors can I adjust if primer-dimers persist after optimizing the temperature? If primer-dimers persist, consider these additional strategies:
- Primer Design: Re-evaluate your primers for self-complementarity and 3'-end complementarity using design tools [7] [22].
- Primer Concentration: Lowering the primer concentration (e.g., from 1µM to 0.1-0.5µM) reduces the chance of primers encountering each other [2] [9].
- Hot-Start Polymerase: Use a hot-start enzyme to prevent polymerase activity during reaction setup at low temperatures, a common period for primer-dimer formation [2] [11].
- Touchdown PCR: Implement a protocol that starts with a higher annealing temperature and gradually decreases it in subsequent cycles, favoring specific product accumulation early on [11].
Experimental Protocol: Using a Temperature Gradient to Find the Optimal Annealing Temperature
1. Objective To empirically determine the annealing temperature that provides the maximum yield of a specific PCR product while minimizing or eliminating non-specific amplification and primer-dimer formation.
2. Background The success of PCR is highly dependent on the annealing temperature (Ta). A Ta that is too low leads to non-specific binding and primer-dimer formation, while a Ta that is too high may reduce or prevent primer binding, resulting in low yield or PCR failure [5]. A gradient thermal cycler is used to test a range of temperatures in a single run, providing a fast and reliable optimization method [5].
3. Materials and Reagents
- DNA template
- Forward and reverse primers
- PCR master mix (containing buffer, dNTPs, and hot-start DNA polymerase)
- Sterile, nuclease-free water
- Gradient thermal cycler
- Equipment for agarose gel electrophoresis
4. Procedure Step 1: Prepare the Master Mix Calculate the volumes required for all reactions plus ~10% extra to account for pipetting error. Combine the following components in a sterile tube on ice:
| Component | Final Concentration | Volume per 25 µL Reaction |
|---|---|---|
| PCR Buffer (10X) | 1X | 2.5 µL |
| dNTP Mix (10 mM) | 200 µM | 0.5 µL |
| Forward Primer (20 µM) | 0.2 µM | 0.25 µL |
| Reverse Primer (20 µM) | 0.2 µM | 0.25 µL |
| DNA Polymerase | 1.25 U | 0.25 µL |
| Template DNA | 10 - 100 ng | Variable |
| Nuclease-free Water | - | To 25 µL |
Mix the master mix by pipetting gently. Aliquot equal volumes into each PCR tube.
Step 2: Set Up the Thermal Cycler Program the thermal cycler with a standard 3-step PCR protocol [9]:
- Initial Denaturation: 94–98°C for 2–5 minutes.
- Amplification Cycles (30–35 cycles):
- Denaturation: 94–98°C for 10–30 seconds.
- Annealing: GRADIENT from 50°C to 70°C for 20–40 seconds.
- Extension: 68–72°C for 1 minute per 1 kb of product.
- Final Extension: 68–72°C for 5–10 minutes.
- Hold: 4–10°C.
Activate the gradient function and assign the desired temperature range to the wells/columns that correspond to your reaction tubes.
Step 3: Analyze Results via Gel Electrophoresis After the run, analyze the PCR products using agarose gel electrophoresis.
- A successful optimization will show a clear, single band of the expected size at one or more of the higher temperatures in the gradient.
- Primer-dimers will appear as a fuzzy smear or fast-migrating band, typically below 100 bp, and will be more prominent at the lower temperatures [2].
- Select the highest annealing temperature that still produces a strong, specific amplicon for all future experiments.
Visual Guide: The Optimization Workflow
The following diagram illustrates the logical process for using a temperature gradient to troubleshoot and eliminate primer-dimer formation.
Research Reagent Solutions
The following table details key reagents and their optimized roles in preventing primer-dimer formation.
| Reagent / Tool | Function in Optimization | Key Considerations for Primer-Dimer Prevention |
|---|---|---|
| Gradient Thermal Cycler | Enables simultaneous testing of multiple annealing temperatures to find the "sweet spot" [5]. | Critical for empirical determination of the highest Ta that provides specific amplification. |
| Hot-Start DNA Polymerase | A modified enzyme inactive until a high-temperature step, preventing activity during reaction setup [2] [11]. | Crucial for minimizing primer-dimer formation that occurs at room temperature before PCR begins. |
| Primer Design Software (e.g., Primer-BLAST, Primer3) | Computes Tm and checks for self-complementarity, hairpins, and dimerization potential [7] [20]. | Select primers with low "self-complementarity" and "self 3'-complementarity" scores [7]. |
| DMSO | An additive that can help denature DNA secondary structures and lower effective Tm [9]. | Can be helpful for GC-rich templates, but requires re-optimization of annealing temperature. |
| SYBR Green Assay | A fluorescent dye that binds double-stranded DNA, used in qPCR [9]. | Essential for testing: Run a no-template control (NTC) to detect primer-dimer amplification as a false-positive signal [10]. |
Primer-dimer formation is a common challenge in polymerase chain reaction (PCR) experiments, often leading to reduced yield of the desired amplicon and compromised data interpretation. This nonspecific amplification occurs when primers anneal to each other instead of the target DNA template, creating short, unintended DNA fragments. This technical guide provides detailed component titration protocols to systematically optimize PCR conditions, minimizing primer-dimer formation while maintaining high amplification efficiency for researchers and drug development professionals.
What is Primer-Dimer and Why Does It Form?
Primer-dimer is a small, unintended DNA fragment that can form during PCR through two primary mechanisms: self-dimerization (a single primer containing complementary regions) or cross-primer dimerization (two primers with complementary regions binding to each other). These structures create free 3' ends that DNA polymerase can extend, consuming reaction resources that would otherwise amplify your target sequence. [2]
The formation of primer-dimer is favored by several factors including poorly designed primers with self-complementary regions, excessive primer concentrations, suboptimal annealing temperatures, and polymerase activity during reaction setup at room temperature. Understanding these mechanisms is crucial for implementing effective optimization strategies. [2] [11]
Troubleshooting Guides
Problem: Excessive Primer-Dimer Formation in Gel Electrophoresis
Issue Description: After PCR amplification, gel electrophoresis shows a fuzzy smear or band below 100 bp in addition to or instead of your target amplicon.
Identification Tips:
- Short length: Primer dimers typically migrate below 100 bp. [2]
- Smeary appearance: They often appear as fuzzy smears rather than well-defined bands. [2]
- No-template control validation: Primer dimers will appear in a no-template control (NTC) reaction since they don't require template DNA. [2]
Solution Approach: Implement a systematic titration of PCR components to favor specific primer-template binding over primer-primer interactions.
Systematic Component Titration Protocol
Primer Concentration Optimization
Objective: Identify the minimum primer concentration that provides robust target amplification without promoting primer-dimer formation.
Background: High primer concentrations increase the likelihood of primer-primer interactions and nonspecific amplification. [46] [13]
Table 1: Primer Concentration Optimization Guide
| Concentration Range (μM) | Expected Outcome | Recommendation |
|---|---|---|
| >1.0 | High risk of primer-dimer and nonspecific products | Avoid except for specialized applications |
| 0.5-1.0 | Standard working range | Suitable for most conventional PCR applications |
| 0.1-0.5 | Optimal for reducing primer-dimer | Recommended starting point for optimization |
| <0.1 | Potential for reduced or failed amplification | May require increased cycle number |
Experimental Protocol:
- Prepare a master mix containing all PCR components except primers.
- Aliquot equal volumes of the master mix into 5 PCR tubes.
- Add primers to achieve these final concentrations: 1.0 μM, 0.5 μM, 0.25 μM, 0.1 μM, and 0.05 μM.
- Include a no-template control for each concentration to specifically monitor primer-dimer formation.
- Run PCR using your standard cycling conditions.
- Analyze results by gel electrophoresis, noting both target amplification efficiency and primer-dimer formation.
Interpretation: Select the lowest primer concentration that provides strong target amplification with minimal primer-dimer. For difficult templates (long amplicons, GC-rich targets), you may need to use concentrations at the higher end of the optimal range (0.3-1.0 μM). [46]
Magnesium Ion (Mg²⁺) Concentration Titration
Objective: Determine the optimal Mg²⁺ concentration that supports efficient polymerase activity while maintaining amplification specificity.
Background: Magnesium ions function as essential cofactors for DNA polymerase activity by enabling dNTP incorporation and stabilizing primer-template binding. However, excessive concentrations reduce specificity and promote mispriming. [46] [47]
Table 2: Magnesium Concentration Optimization Guide
| Mg²⁺ Concentration (mM) | Expected Outcome | Recommendation |
|---|---|---|
| <1.5 | Potential PCR failure or weak amplification | Increase concentration |
| 1.5-2.0 | Optimal for most applications with Taq polymerase | Standard working range [47] |
| 2.0-4.0 | May increase yield but risk nonspecific products | Titrate carefully |
| >4.0 | High risk of nonspecific amplification and primer-dimer | Generally avoid |
Experimental Protocol:
- Prepare a master mix containing all components except Mg²⁺.
- Aliquot equal volumes into 6-8 PCR tubes.
- Add MgCl₂ or MgSO₄ (depending on polymerase preference) to achieve a concentration series: 0.5, 1.0, 1.5, 2.0, 2.5, 3.0, 4.0 mM.
- Include appropriate positive and negative controls.
- Perform PCR amplification and analyze by gel electrophoresis.
Interpretation: The optimal Mg²⁺ concentration depends on your specific template, primers, and buffer composition. Note that dNTPs chelate Mg²⁺, so the effective concentration is approximately 0.5-1.0 mM less than the added concentration due to dNTP binding. [46] Pfu DNA polymerase often works better with MgSO₄ than with MgCl₂. [13]
DNA Polymerase Concentration and Selection
Objective: Optimize polymerase type and concentration to minimize nonspecific amplification during reaction setup and early PCR cycles.
Background: DNA polymerase concentration affects both yield and specificity. Excessive enzyme can amplify nonspecific products, while insufficient enzyme results in poor yield. [46] [48]
Table 3: DNA Polymerase Selection and Concentration Guide
| Polymerase Type | Key Characteristics | Recommended Concentration | Effect on Primer-Dimer |
|---|---|---|---|
| Standard Taq | Moderate fidelity, no proofreading | 0.5-2.0 units/50 μL reaction [47] | Higher risk without careful setup |
| Hot-Start Taq | Activated at high temperature, reduces pre-PCR activity | 1.0-1.25 units/50 μL reaction | Significantly reduces primer-dimer [2] [48] |
| High-Fidelity | Proofreading activity, high accuracy | Manufacturer's recommendation | Reduces mispriming events |
Experimental Protocol:
- Select 2-3 different polymerase types (e.g., standard, hot-start, high-fidelity).
- For each type, prepare a dilution series: 0.25x, 0.5x, 1x, 2x the manufacturer's recommended concentration.
- Set up reactions on ice to minimize pre-PCR activity for non-hot-start enzymes.
- Include a room-temperature setup control to assess the benefit of hot-start enzymes.
- Run PCR and evaluate specificity and yield.
Interpretation: Hot-start DNA polymerases are particularly effective for reducing primer-dimer as they remain inactive until the first denaturation step, preventing extension of primer-dimers formed during reaction setup. [2] [48] For standard polymerases, setting up reactions on ice and adding polymerase last can help reduce nonspecific amplification. [13]
Comprehensive Workflow for PCR Optimization
The following diagram illustrates the systematic approach to optimizing PCR components to minimize primer-dimer formation:
Frequently Asked Questions (FAQs)
Q1: Why do I still see primer-dimer even after using hot-start polymerase? Hot-start polymerases reduce but don't completely eliminate primer-dimer formation. They prevent enzyme activity during reaction setup, but primer-dimer can still form during subsequent cycles if primers have significant complementarity or annealing temperatures are too low. Consider revisiting your primer design and optimizing annealing temperature. [2] [48]
Q2: How can I distinguish primer-dimer from my target amplicon on a gel? Primer-dimers have two telltale characteristics: (1) They're short (typically below 100 bp) and (2) They often appear as fuzzy smears rather than sharp, well-defined bands. Running your gel longer can help separate primer-dimers from your target product. [2]
Q3: What is the ideal primer-to-template ratio for minimizing primer-dimer? There's no universal ideal ratio, but lowering primer concentration while maintaining sufficient template is generally beneficial. Aim for primer concentrations between 0.1-0.5 μM and ensure adequate template (10^4-10^7 copies for genomic DNA). The key is achieving a lower primer-to-template ratio to favor specific binding. [2] [47]
Q4: Can I completely eliminate primer-dimer from my PCR reactions? In many cases, primer-dimer can be reduced to undetectable levels, but complete elimination may not always be possible or necessary. The goal is to minimize it sufficiently so it doesn't interfere with your application. Some applications like qPCR are more sensitive to primer-dimer than conventional PCR followed by gel electrophoresis. [2]
Q5: How does annealing temperature affect primer-dimer formation? Higher annealing temperatures increase stringency, favoring specific primer-template binding over primer-primer interactions. If you're seeing primer-dimer, try increasing the annealing temperature in 2-3°C increments, up to 5°C below the primer Tm. Using a gradient thermal cycler can help identify the optimal temperature efficiently. [2] [13]
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for PCR Optimization and Primer-Dimer Reduction
| Reagent | Function | Optimization Guidelines |
|---|---|---|
| Hot-Start DNA Polymerase | Reduces nonspecific amplification during reaction setup; remains inactive until high-temperature activation | Choose antibody-inactivated versions for true hot-start capability; follow manufacturer's activation guidelines [48] |
| dNTP Mix | Building blocks for DNA synthesis | Use 200 μM of each dNTP as starting point; lower concentrations (50-100 μM) can enhance fidelity but may reduce yield [47] |
| Magnesium Salts | Cofactor for polymerase activity | Titrate between 1.5-4.0 mM; note that MgCl₂ works well with Taq, while MgSO₄ may be better for proofreading enzymes [13] |
| PCR Enhancers | Improve amplification of difficult templates | Use DMSO (1-10%), formamide (1.25-10%), or Betaine (0.5-2.5 M) for GC-rich targets; may require re-optimization of annealing temperature [20] |
| Nuclease-Free Water | Reaction solvent | Ensure purity and absence of nucleases; use for all reagent preparations and dilutions |
Advanced Techniques and Future Directions
For particularly challenging applications, consider these advanced approaches:
Touchdown PCR: Gradually decreasing the annealing temperature over cycles can increase specificity in early cycles when primer-dimer is most likely to form. [13]
SAMRS Technology: Self-Avoiding Molecular Recognition Systems incorporate modified bases that pair with natural DNA but not with other SAMRS bases, fundamentally preventing primer-primer interactions. [6]
Additive Optimization: For GC-rich templates or complex secondary structures, additives like DMSO (1-10%), formamide (1.25-10%), or betaine (0.5-2.5 M) can help denature templates and improve specificity, though they may require additional optimization of other parameters. [20] [13]
Systematic titration of PCR components—particularly primers, magnesium ions, and DNA polymerase—provides a robust framework for minimizing primer-dimer formation while maintaining amplification efficiency. The protocols outlined in this guide emphasize empirical testing with controlled concentration gradients, enabling researchers to establish optimal conditions for their specific experimental systems. Remember that primer-dimer is not necessarily a sign of a flawed experiment, but rather a common challenge that can be systematically addressed through careful optimization. [2]
Troubleshooting Guides and FAQs
Frequently Asked Questions
Q1: How do DMSO, BSA, and Betaine specifically help in reducing primer-dimer formation and non-specific amplification?
Primer-dimer formation and non-specific amplification often occur due to low-stringency conditions where primers anneal to non-target sequences or to each other. The strategic use of additives enhances specificity through distinct mechanisms:
- Betaine (1-1.7 M) acts as a universal PCR enhancer by reducing the dependence of DNA melting on base pair composition. It equalizes the melting temperatures of GC-rich and AT-rich regions, which promotes uniform primer binding to the true template and discourages mispriming events that lead to primer-dimers and non-specific products [49] [50].
- DMSO (2-10%) primarily disrupts the secondary structures in both the DNA template and primers. By reducing the stability of these structures, it prevents the polymerase from stalling and minimizes chances for primers to form dimers or bind to non-target sites. Note that it can also slightly reduce Taq polymerase activity [49] [51].
- BSA (0.1-0.8 mg/ml) functions as a scavenger. It binds to inhibitors commonly found in sample preparations (e.g., phenolic compounds or salts) that can otherwise force the reaction into low-stringency conditions, thereby indirectly promoting specificity and protecting the polymerase [49] [52].
Q2: My PCR target is extremely GC-rich (>80%). A single additive isn't working. What should I do?
For extremely challenging GC-rich templates, a combinatorial approach is often necessary. A powerful validated mixture includes:
- Final Concentrations: 1-1.7 M Betaine, 5% DMSO, and 50 µM 7-deaza-dGTP (a dGTP analog) [51].
- Mechanism: This combination tackles the problem synergistically. Betaine homogenizes DNA melting, DMSO disrupts persistent secondary structures, and 7-deaza-dGTP incorporates into the newly synthesized DNA, preventing the formation of tight secondary structures without compromising base pairing [51] [53]. This specific cocktail has been proven essential for amplifying DNA sequences with GC content ranging from 67% to 79% [51].
Q3: At what point should I add these additives to my PCR reaction, and do they affect the thermal cycling conditions?
Additives should be included in the initial reaction setup. For the most challenging templates, a controlled heat denaturation step before cycling can be highly beneficial:
- Protocol: Combine your DNA template and primer in a low-salt buffer (e.g., 10 mM Tris-HCl, pH 8.0). Heat this mixture to 98°C for 5 minutes, then cool on ice. After this, add the rest of the PCR components, including the master mix with your chosen additives [54].
- Thermal Cycling: Standard thermal cycling parameters are usually sufficient. However, because additives like DMSO lower the DNA melting temperature (Tm), you might consider empirically lowering the annealing temperature by 1-3°C for optimal results [49] [55].
Troubleshooting Common Problems
Problem: The PCR yield is low, and non-specific bands are present.
- Potential Cause: The concentration of DMSO might be too high, inhibiting the polymerase, or the Mg²⁺ concentration might not be optimized.
- Solution: Titrate the DMSO concentration (test between 2-10%) and optimize the Mg²⁺ concentration (e.g., from 1.0 to 4.0 mM in 0.5 mM intervals) [49]. Ensure you are using a high-quality DNA template.
Problem: There is no amplification product at all.
- Potential Cause: The template may contain potent inhibitors, or the secondary structures are too stable.
- Solution: Increase the concentration of BSA (up to 0.8 mg/ml) to neutralize inhibitors [49] [52]. Implement the pre-PCR heat denaturation step in low-salt buffer [54]. Consider using a polymerase mixture or a specialized high-performance polymerase designed for difficult templates.
Table 1: Working Concentrations and Mechanisms of Key PCR Additives
| Additive | Common Working Concentration | Primary Mechanism of Action | Key Considerations |
|---|---|---|---|
| DMSO | 2 - 10% | Disrupts DNA secondary structure; lowers melting temperature (Tm) [49] [56]. | High concentrations can inhibit polymerase activity; requires titration [49]. |
| Betaine | 1.0 - 1.7 M | Equalizes the melting temperature of DNA; reduces base-composition dependence [49] [51]. | Use betaine or betaine monohydrate; betaine hydrochloride may affect pH [49]. |
| BSA | 0.1 - 0.8 mg/ml | Binds and neutralizes PCR inhibitors (e.g., phenols, humic acids) [49] [52]. | Particularly useful for contaminated or "dirty" samples; minimal effect on clean DNA [52]. |
| 7-deaza-dGTP | 50 µM (as partial substitute for dGTP) | dGTP analog that prevents formation of stable secondary structures [51]. | Often used in a 1:3 ratio with dGTP; effective for extreme GC-rich targets [51] [56]. |
Table 2: Additive Cocktails for Specific Template Challenges
| Template Challenge | Recommended Additive Cocktail | Reported Efficacy |
|---|---|---|
| General GC-rich (60-70% GC) | 5% DMSO + 1 M Betaine [51] [56] | Improves yield and specificity for many difficult amplicons. |
| Extreme GC-rich (>70% GC) or Strong Secondary Structures | 5% DMSO + 1.3 M Betaine + 50 µM 7-deaza-dGTP [51] | Essential for specific amplification of sequences with GC content up to 79% [51]. |
| GC-rich Templates with Potential Inhibitors | 5% DMSO + 0.8 mg/ml BSA [52] | BSA acts as a co-enhancer, significantly boosting yield in the presence of solvents [52]. |
Experimental Protocol: Amplification of a GC-Rich Target
This protocol is adapted from a published study that successfully amplified GC-rich disease genes using a powerful additive mixture [51].
Objective: To amplify a GC-rich DNA target (e.g., 67-79% GC content) that has proven refractory to standard PCR conditions.
Reagents and Materials:
- Template DNA (e.g., 100 ng genomic DNA)
- Forward and Reverse Primers
- Taq DNA Polymerase with recommended 10x Buffer
- dNTP Mix (including 7-deaza-dGTP)
- MgCl₂ (25-50 mM stock)
- Betaine (5 M stock solution)
- DMSO
- Nuclease-free Water
Procedure:
- Prepare the PCR Master Mix on ice, in the following order:
- Nuclease-free Water: to a final volume of 25 µL
- 10x PCR Buffer: 1x final concentration
- MgCl₂: 2.5 mM final concentration (optimize if needed)
- dNTP Mix: 200 µM each dATP, dCTP, dTTP; 150 µM dGTP
- 7-deaza-dGTP: 50 µM final concentration
- Betaine: 1.3 M final concentration (from 5M stock)
- DMSO: 5% (v/v) final concentration
- Forward and Reverse Primers: 10 pmol each
- Template DNA: 100 ng
- Taq DNA Polymerase: 1.25 units
Thermal Cycling:
- Initial Denaturation: 94°C for 5 minutes
- 30-40 Cycles of:
- Denaturation: 94°C for 30 seconds
- Annealing: 60°C for 30 seconds (optimize based on primer Tm)
- Extension: 72°C for 45 seconds per kb
- Final Extension: 72°C for 5 minutes
- Hold: 4°C
Analysis: Analyze 5 µL of the PCR product by agarose gel electrophoresis.
Workflow Visualization
The following diagram illustrates the logical workflow for troubleshooting a challenging PCR experiment using the additives discussed.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for PCR Enhancement Experiments
| Reagent / Material | Function in the Protocol | Notes for Researchers |
|---|---|---|
| Betaine (Monohydrate) | Equalizes DNA melting temperatures; crucial for GC-rich targets [49] [51]. | Prepare a 5M stock solution in nuclease-free water. Avoid betaine-HCl as it can alter pH [49]. |
| DMSO (Molecular Biology Grade) | Disrupts DNA secondary structures like hairpins and stem-loops [49] [56]. | Always use high-purity grade. Titrate for each application as it can inhibit polymerase at high levels [49]. |
| Molecular Grade BSA | Neutralizes a wide range of PCR inhibitors present in biological samples [49] [52]. | Acetylated BSA is often preferred for its stability and low enzyme activity. |
| 7-deaza-2'-deoxyguanosine (7-deaza-dGTP) | dGTP analog that reduces hydrogen bonding, easing amplification through GC-rich regions [51] [56]. | Typically used as a partial substitute for dGTP (e.g., 50 µM 7-deaza-dGTP with 150 µM dGTP) [51]. |
| Thermostable DNA Polymerase | Enzymatic engine for DNA synthesis. | For long or complex templates, consider polymerases with proofreading activity or specialized enzyme blends [50] [55]. |
FAQ: Preventing Primer-Dimer Formation
Why is reaction assembly on ice crucial for preventing primer-dimer?
A: Assembling your PCR reaction on ice is a fundamental practice to prevent non-specific interactions like primer-dimer formation before thermal cycling begins. When reagents are mixed at room temperature, the DNA polymerase can retain some enzymatic activity. This allows primers that have transiently bound to each other to be extended, creating short, unwanted primer-dimer products [13]. Keeping the reaction mixture cold on ice suppresses this low-level activity until a proper denaturation step occurs in the thermal cycler. For best results, use a hot-start DNA polymerase, which is inactive until heated to a high temperature, providing a double layer of protection [2] [13].
What is the connection between immediate thermal cycling and primer-dimer reduction?
A: Immediate thermal cycling ensures that once the reaction tube is transferred from your ice bath to the pre-heated thermal cycler, the reaction is rapidly raised to the denaturation temperature (typically 94–98°C). This minimizes the time window where primers could anneal to each other at intermediate, non-specific temperatures. A delay at room temperature after assembly allows more opportunity for primers to bind nonspecifically, which can then be extended by any residual polymerase activity, leading to primer-dimer formation [13].
My primers still form dimers even on ice. What else can I optimize?
A: While assembly on ice and immediate cycling are critical, other factors contribute to primer-dimer formation. Consider these optimizations:
- Primer Design: Check that your primers, especially at their 3' ends, do not have complementary sequences to each other. Using primer design software is highly recommended [2] [8] [57].
- Annealing Temperature: Increase the annealing temperature in increments of 2–3°C. A higher temperature favors specific primer-template binding and discourages the weaker binding between primers [2] [58] [13].
- Primer Concentration: High primer concentrations increase the chance of primers meeting and dimerizing. Test lower primer concentrations within the standard range of 0.1–1 µM [2] [8] [57].
- Mg²⁺ Concentration: Excess Mg²⁺ can reduce reaction specificity and promote non-specific amplification. Ensure the Mg²⁺ concentration is optimized for your specific polymerase and reaction [13] [59].
Troubleshooting Guide: Primer-Dimer Formation
| Observation | Potential Cause | Recommended Solution |
|---|---|---|
| Fuzzy band/smear below 100 bp on gel | Nonspecific primer annealing and extension at low temperatures | Use hot-start DNA polymerase; assemble reactions on ice; increase annealing temperature [2] [13] |
| Primer-dimer in No-Template Control (NTC) | Primers annealing to each other, not template-dependent | Redesign primers to avoid 3' complementarity; lower primer concentration; use a hot-start polymerase [2] [8] |
| Weak desired product band with strong primer-dimer | Primer-dimer formation outcompeting target amplification | Increase annealing temperature; verify template quality and quantity; switch to a two-step PCR protocol [58] [13] [59] |
| Persistent primer-dimer after optimization | Suboptimal primer design with high self-complementarity | Redesign primers using validated software tools; consider nested PCR for difficult targets [13] [59] |
Experimental Protocol: Low-Temperature Reaction Assembly
Objective: To assemble a PCR reaction mixture in a manner that minimizes primer-dimer formation by maintaining low temperatures and using a hot-start DNA polymerase.
Materials:
- PCR reagents (DNA polymerase, buffer, dNTPs, MgCl₂, primers, template DNA, nuclease-free water)
- Ice bucket with crushed ice
- Sterile, nuclease-free microcentrifuge tubes and pipette tips
- Thermal cycler pre-heated to initial denaturation temperature (e.g., 95°C)
Methodology:
- Preparation: Thaw all PCR reagents (except the template and polymerase, if recommended) and quickly place them on ice. Centrifuge briefly to gather the liquid at the bottom of the tubes.
- Master Mix Assembly on Ice: On ice, prepare a master mix in the following order to ensure homogeneity:
- Aliquoting and Adding Template: Gently mix the master mix by pipetting up and down. Aliquot the appropriate volume into individual PCR tubes placed on ice. Lastly, add the template DNA to each tube, pipetting to mix. Keep tubes on ice until ready to load the thermal cycler.
- Immediate Thermal Cycling: Transfer the PCR tubes directly from ice to the thermal cycler, which has been pre-heated to the initial denaturation temperature (e.g., 95°C for 2–5 minutes). Start the cycling program immediately. [13]
Quantitative Data for PCR Optimization
Table 1: Optimized Thermal Cycling Parameters to Minimize Primer-Dimer
| Cycling Step | Temperature Range | Time Duration | Special Considerations |
|---|---|---|---|
| Initial Denaturation | 94–98°C | 1–3 minutes | Critical for hot-start polymerase activation and full DNA denaturation [58] |
| Denaturation | 94–98°C | 15–30 seconds | Longer times may be needed for GC-rich templates [58] [13] |
| Annealing | Tm +3°C to Tm -5°C | 15–30 seconds | Optimize empirically; use a gradient cycler for best results [58] [57] |
| Extension | 68–72°C | 15–60 sec/kb | Time depends on polymerase speed and amplicon length [58] [57] |
| Cycle Number | 25–35 cycles | - | Avoid over-cycling (>45 cycles) to prevent by-product accumulation [58] [13] |
| Final Extension | 68–72°C | 5–15 minutes | Ensures full-length products are synthesized [58] |
Table 2: Key Research Reagent Solutions for Primer-Dimer Prevention
| Reagent | Function in Prevention | Optimization Notes |
|---|---|---|
| Hot-Start DNA Polymerase | Inactive at room temperature; prevents pre-cycling extension of primers | The cornerstone reagent; always prefer over standard polymerases [2] [13] |
| Ultrapure dNTPs | Balanced substrates for DNA synthesis | Use at 200 µM each; unbalanced concentrations can increase error rate [57] [59] |
| Magnesium Salt (MgCl₂/MgSO₄) | Essential cofactor for polymerase activity | Titrate concentration (often 1.5-2.0 mM); excess promotes non-specific binding [13] [57] |
| Optimized PCR Buffer | Provides optimal ionic and pH conditions | Some buffers allow for a universal annealing temperature (~60°C), simplifying optimization [58] |
| PCR Additives (e.g., DMSO, Betaine) | Aid in denaturing complex templates (GC-rich) | Can help reduce secondary structures that might favor mis-priming; use at recommended concentrations [58] [13] |
Workflow and Decision Diagrams
PCR Assembly on Ice Workflow
Primer-Dimer Troubleshooting Path
Validation Frameworks and Technology Comparisons for Primer-Dimer Management
In polymerase chain reaction (PCR) research, method validation is the cornerstone of data integrity and reliable results. For researchers and drug development professionals, establishing robust PCR methods is essential for generating actionable data. A pervasive challenge in this process is the formation of primer dimers (PDs), which are short, unintended DNA fragments that form when primers anneal to each other instead of the target DNA template [1] [2]. Primer dimers compete for PCR reagents, can inhibit amplification of the desired target sequence, and in quantitative PCR (qPCR), they interfere with accurate quantification, thereby compromising the specificity, sensitivity, and robustness of an assay [1] [60]. This technical guide, framed within the broader thesis of reducing primer-dimer formation, provides troubleshooting FAQs and detailed protocols to fortify your method validation.
Frequently Asked Questions (FAQs) on Primer Dimers
Q1: What exactly is a primer dimer and how does it impact assay sensitivity and specificity?
A primer dimer is a small, double-stranded DNA fragment, typically between 30-100 base pairs, formed when two PCR primers anneal to each other via complementary base pairs and are extended by the DNA polymerase [1] [2]. Its impact is twofold:
- Sensitivity: PDs consume primers, nucleotides, and enzyme, reducing the reagents available for amplifying the true target. This can lead to a decrease in the final yield of the desired product and a higher, less sensitive limit of detection (LOD) [1] [11].
- Specificity: PDs are non-specific amplification products. In qPCR using intercalating dyes like SYBR Green, they generate false fluorescence signals, leading to inaccurate cycle threshold (Ct) values and potentially masking the true result [60].
Q2: During method validation, how can I conclusively distinguish primer dimer from my target amplicon?
You can distinguish them using the following methods:
- Gel Electrophoresis: After PCR, PDs typically appear as a diffuse smear or a sharp band below 100 bp, which is distinguishable from the larger, well-defined band of the target amplicon. Running the gel longer can help separate these small fragments [2].
- Melting Curve Analysis: In qPCR with intercalating dyes, the PDs, being shorter and with a different sequence, will melt at a lower temperature than the target amplicon. A specific target will show a single, sharp peak, while the presence of PDs will manifest as an additional peak or a broad "waveform" around 70°C [1] [60].
- No-Template Control (NTC): This is a critical validation control. A reaction where template DNA is replaced with water should not amplify the target. If a band or signal appears in the NTC, it is almost certainly due to primer dimer or contamination, confirming the assay's specificity is compromised [2] [60].
Q3: My assay is for diagnostic use. What is the difference between core and full process validation for ensuring primer-dimer-free results?
The choice depends on the intended use and regulatory requirements [61]:
- Core Validation focuses on the essential analytical components of the PCR assay itself. It evaluates parameters like specificity, sensitivity (LOD), and linearity under controlled conditions. This is ideal for early-stage research, assay optimization, or Research Use Only (RUO) applications.
- Full Process Validation is essential when results inform clinical decisions or are submitted to regulators. It encompasses the entire workflow—from sample extraction and preparation to data analysis—ensuring every step is controlled and robust against variables that could induce primer dimer formation [61].
Troubleshooting Guide: Reducing Primer-Dimer Formation
The following table summarizes the primary causes of primer dimer formation and evidence-based solutions to address them.
| Cause of Primer Dimer | Troubleshooting Strategy | Specific Experimental Protocol / Note |
|---|---|---|
| Complementary primer sequences, especially at the 3' ends [1] [20] | Optimize primer design using software (e.g., Primer-BLAST, Primer3) to avoid self-complementarity and cross-complementarity. | Design primers 18-30 bases long with 40-60% GC content. Ensure the 3' ends do not contain G or C runs and have no more than 2-3 complementary bases with the other primer [20] [5] [8]. |
| Low annealing temperature [60] [5] | Increase the annealing temperature to promote specific binding. | Perform a gradient PCR (e.g., testing from 55°C to 68°C) to determine the highest temperature that yields specific product without dimers [5]. |
| High primer concentration [60] [8] | Lower the concentration of primers in the reaction. | Test a primer concentration gradient, typically from 0.1 to 0.5 µM, to find the lowest concentration that supports efficient amplification [8]. |
| Polymerase activity at low temperatures during reaction setup [1] [5] | Use a hot-start DNA polymerase. | These enzymes are inactive until a high-temperature activation step (e.g., 95°C), preventing low-temperature mis-priming and extension [1] [2]. |
| Low template concentration or quality [60] | Ensure an adequate amount of high-quality, pure template DNA is used. | Optimize template concentration. Use controls to verify template quality and avoid impurities that can inhibit amplification and promote dimer formation [60]. |
| Excessive PCR cycle numbers [60] [5] | Reduce the number of amplification cycles. | For most applications, 30-35 cycles are sufficient. Excess cycles can amplify low-level primer dimers formed in early cycles [5]. |
Advanced Techniques for Intractable Cases
For persistent primer dimer issues, consider these advanced strategies:
- Hot-Start PCR Methods: Beyond standard hot-start enzymes, methods include wax barriers, slow release of magnesium, or antibodies that inhibit the polymerase until the first denaturation step [1].
- Primer Structural Modifications: Techniques like HANDS (Homo-Tag Assisted Non-Dimer System) add a complementary tail to the 5' end, forming a stem-loop that prevents dimerization [1].
- Sequence-Specific Probes: In qPCR, switching from SYBR Green to TaqMan probes or molecular beacons ensures that fluorescence is generated only upon hybridization to the specific target sequence, preventing signal acquisition from any PDs that form [1].
Experimental Protocols for Validation
Protocol 1: Establishing Specificity via Melting Curve Analysis
Purpose: To validate that the amplification signal is derived from the specific target amplicon and not from primer dimers in a SYBR Green qPCR assay [1] [60].
Methodology:
- Run qPCR: Perform the qPCR run with your optimized protocol and primers.
- Initiate Melting Curve: After the final amplification cycle, the instrument slowly increases the temperature from approximately 60°C to 95°C while continuously monitoring fluorescence.
- Analyze Data: Plot the negative derivative of fluorescence over temperature (-dF/dT) versus temperature. A specific reaction will produce a single, sharp peak at the melting temperature (Tm) of the target amplicon. A broader peak or a separate peak at a lower temperature (~70-75°C) indicates the presence of primer dimers.
Protocol 2: Determining Robustness by Testing Annealing Temperature
Purpose: To demonstrate that the PCR assay remains specific and efficient under small, intentional variations in a critical parameter (annealing temperature) [5].
Methodology:
- Set Up Reactions: Prepare a master mix containing all PCR components and aliquot it into several tubes.
- Create Gradient: Using a thermal cycler with a gradient function, set a range of annealing temperatures (e.g., 5°C above and below the calculated Tm).
- Run PCR and Analyze: Execute the PCR protocol and analyze the products via gel electrophoresis. The optimal and robust annealing temperature is the highest temperature that produces a strong, specific target band with minimal to no primer dimer smear.
The workflow below outlines the key decision points for selecting the right validation path for your PCR assay.
Diagram 1: A workflow for choosing between core and full process validation for your PCR assay, based on its intended use [61].
The Scientist's Toolkit: Key Reagents for Validation
The following table details essential reagents used in validating and optimizing PCR assays to minimize primer-dimer formation.
| Reagent / Material | Function in Validation & Primer Dimer Prevention |
|---|---|
| Hot-Start DNA Polymerase | A modified enzyme inactive at room temperature, preventing enzymatic extension of primed dimers during reaction setup. It is activated by a high-temperature step, drastically improving specificity [1] [2]. |
| HPLC-Purified Primers | High-purity primers ensure the correct sequence is dominant, reducing short primers and truncated sequences that are more prone to form dimers [5]. |
| SYBR Green I Dye | A nonspecific intercalating dye used for qPCR and melting curve analysis. It is crucial for detecting and diagnosing the presence of primer dimers through their characteristic lower melting temperature [1] [60]. |
| Sequence-Specific Probes (e.g., TaqMan) | These probes provide a target-specific signal in qPCR. Since fluorescence requires probe binding and cleavage, they prevent false-positive signals from primer dimers, enhancing specificity validation [1]. |
| No-Template Control (NTC) | A control reaction containing all PCR components except the template DNA. The presence of amplification in the NTC is a direct indicator of primer-dimer formation or contamination [2] [60]. |
Digital PCR (dPCR) is a powerful nucleic acid quantification technology that provides absolute quantification without the need for a standard curve. By partitioning a PCR reaction into tens of thousands of individual reactions, dPCR enables precise measurement of target sequences based on Poisson statistics. This technique offers significant advantages for applications requiring high precision, including copy number variation analysis, rare mutation detection, and genetically modified organism (GMO) quantification. Within the context of reducing primer-dimer formation, dPCR's partitioning nature inherently minimizes the impact of such nonspecific amplification products on quantification accuracy, as primer-dimers are typically confined to a subset of partitions rather than affecting the entire reaction.
Core Principles and Advantages
- Absolute Quantification: dPCR provides direct, absolute quantification of nucleic acid targets without requiring standard curves, measuring results in copies per microliter [62].
- Partitioning Technology: Reactions are subdivided into numerous partitions; target molecules are randomly distributed, with some partitions containing one or more molecules and others containing none [62].
- Poisson Statistics: Quantification uses Poisson statistical models to calculate target concentration based on the proportion of positive and negative partitions [62].
- Enhanced Precision: Partitioning increases quantification precision, with optimal performance achieved when the average number of copies per partition is between 0.5 to 3 [63] [62].
- Superior Multiplexing: dPCR enables precise measurement of more than two targets in a single reaction, known as 'higher order multiplexing' [62].
- Reduced Inhibitor Effects: dPCR is less prone to PCR inhibitors compared to traditional qPCR, making it suitable for complex samples [64].
Troubleshooting Guides and FAQs
Assay Design and Optimization
Q: What are the critical factors in dPCR primer and probe design?
A successful dPCR assay requires careful attention to several design parameters:
- Complementarity: Ensure primers have no self- or inter-complementarity to prevent primer-dimer formation [63].
- Amplicon Length: Keep amplicons short, especially for degraded samples like FFPE DNA or cfDNA [63].
- Melting Temperature: Design primers with appropriate and consistent melting temperatures [65].
- Concentration Optimization: Use primer concentrations between 0.5-0.9 μM and probe concentrations at approximately 0.25 μM per reaction for optimal fluorescence intensity [63].
- Storage Conditions: Store lyophilized primers and probes in TE buffer (pH 8.0 for most probes, pH 7.0 for Cy5 and Cy5.5-labeled probes) to prevent degradation [63].
Q: How can I minimize primer-dimer formation in dPCR assays?
Primer-dimer formation can significantly impact dPCR accuracy, particularly when using DNA-binding dyes. Implement these strategies:
- Hot-Start Polymerases: Use polymerases that remain inactive until a high-temperature activation step to prevent nonspecific amplification during reaction setup [11] [66].
- Careful Primer Design: Utilize primer design software to avoid complementary regions within and between primers [11].
- Optimized Annealing Temperatures: Increase annealing temperatures or use touchdown PCR to promote specific binding [11] [66].
- Reduced Primer Concentration: Lower primer concentrations within the optimal range of 0.1-1 μM to minimize primer-dimer formation [13].
Sample Preparation and Quality Control
Q: What sample quality considerations are critical for dPCR?
Sample purity and integrity significantly impact dPCR results:
- Purity Requirements: Contaminants including alcohols, salts, humic acids, nucleases, urea, phenol, and acidic polysaccharides can impair enzyme activity, fluorescence detection, and amplification efficiency [63].
- Integrity Assessment: For strongly degraded templates (FFPE DNA, cfDNA), use shorter amplicons and consider dedicated extraction kits to recover high-quality DNA [63].
- Input Amount: Ensure the average number of target copies per partition falls within the optimal 0.5-3 range for accurate quantification [63].
- Inhibition Testing: Perform serial dilutions of DNA samples to check for inhibition; measured copies should scale proportionally with dilution [64].
Q: When should I use restriction digestion prior to dPCR?
Restriction digestion is recommended in these specific scenarios:
- Highly Viscous Solutions: Reduces viscosity to enable accurate measurement of higher DNA concentrations [63].
- Linked Gene Copies: Physically separates gene copies to ensure independent segregation into partitions [63].
- Supercoiled Plasmids: Linearizes plasmids to improve primer/probe binding accessibility [63].
- Large DNA Molecules: Fragments templates >30 kb to prevent uneven partitioning and over-quantification [63].
Table 1: Troubleshooting Common dPCR Issues
| Problem | Possible Causes | Solutions |
|---|---|---|
| Poor Partition Separation | Suboptimal probe design, fluorescent crosstalk | Avoid reporter-quencher combinations with overlapping emissions [63] |
| Low Amplitude Signal | Insufficient primer/probe concentration | Increase primer concentration to 0.5-0.9 μM and probes to 0.25 μM [63] |
| Uneven Amplification in Multiplex | Primer interference, differing Tm values | Design primers with Tms within 5°C of each other [66] |
| Inaccurate Quantification | Too many targets/partition | Dilute sample to achieve 0.5-3 copies/partition [63] |
Multiplexing Challenges
Q: What strategies improve multiplex dPCR performance?
Successful multiplexing in dPCR requires careful optimization:
- Primer Compatibility: Validate each primer set individually before multiplexing to ensure specificity and efficiency [67] [66].
- Fluorophore Selection: Use non-overlapping fluorophores with minimal spectral crosstalk [67].
- Balanced Amplification: Design all primer pairs to have similar Tms (within 5°C) to prevent preferential amplification [66].
- Hot-Start Enzymes: Employ hot-start DNA polymerases to minimize primer-dimer formation and improve specificity in complex reactions [66].
Q: What types of duplex assays can be developed in dPCR?
There are two primary configurations for duplex dPCR assays:
- Non-Competing Duplex: Uses two primer pairs targeting different regions; produces four distinct clusters (double-negative, two single-positive, and double-positive) in 2D plots [62].
- Competing Duplex: Employs one primer pair with two probes binding the same region; ideal for SNP detection and rare variant identification [62].
Data Analysis and Interpretation
Q: How do I ensure my dPCR data is in the "digital range"?
The digital range is critical for accurate quantification:
- Partition Analysis: Ensure sufficient partitions contain no template molecules (negative partitions) for proper Poisson statistics [68] [62].
- Optimal Loading: Target approximately 0.5-3 copies per partition to maintain statistical validity [63].
- Threshold Setting: Manually adjust fluorescence thresholds if automatic settings don't properly separate positive and negative partitions [68] [62].
Q: How is copy number calculation performed in dPCR?
dPCR uses Poisson statistics for absolute quantification:
- Fundamental Equation: λ = -ln(1 - k/n), where λ is the average number of targets per partition, n is the total number of partitions, and k is the number of positive partitions [62].
- Alternative Calculation: λ = ln(n) - ln(w), where w is the number of negative partitions [62].
- Concentration Calculation: Copies/μL = (λ × total partitions) / reaction volume [64].
Table 2: Digital PCR Performance Parameters for GMO Quantification
| Parameter | MON-04032-6 Soybean | MON89788 Soybean | Acceptance Criteria |
|---|---|---|---|
| Dynamic Range | 0.05% - 10% GM | 0.1% - 10% GM | Linear across range [64] |
| Linearity | R² > 0.99 | R² > 0.99 | Meets validation criteria [64] |
| Accuracy (Trueness) | Within acceptance | Within acceptance | Comparable to qPCR [64] |
| Precision | Within acceptance | Within acceptance | Meets validation criteria [64] |
| Platform Equivalence | QIAcuity vs. QX200 | QIAcuity vs. QX200 | Equivalent performance [64] |
Experimental Protocols
Protocol 1: Duplex dPCR Assay for GMO Quantification
This protocol, adapted from a study comparing dPCR platforms, enables simultaneous quantification of transgenic and reference genes [64]:
- DNA Extraction: Extract DNA using a validated method (e.g., CTAB buffer or commercial kits).
- DNA Quantification: Measure DNA concentration and purity; perform inhibition testing with serial dilutions.
- Reaction Setup:
- Prepare master mix with 0.5-0.9 μM final primer concentration and 0.25 μM probe concentration.
- Use hot-start DNA polymerase for enhanced specificity.
- Add 1-100 ng DNA template per reaction.
- Partitioning: Load reactions into nanoplates or droplet generation cartridges depending on platform.
- Thermal Cycling:
- Initial activation: 95°C for 10 minutes
- 40 cycles of: 95°C for 30 seconds (denaturation), 60°C for 60 seconds (annealing/extension)
- Final hold: 4°C
- Data Analysis: Use platform-specific software to calculate copies/μL for both targets.
Protocol 2: Restriction Digestion for Complex Templates
For templates requiring restriction digestion prior to dPCR [63]:
- Enzyme Selection: Choose enzymes that don't cut within the amplicon sequence.
- Digestion Reaction: Incubate 100 ng-1 μg DNA with restriction enzymes in appropriate buffer for 1-2 hours.
- Enzyme Inactivation: Heat-inactivate enzymes according to manufacturer recommendations.
- dPCR Setup: Use digested DNA directly in dPCR reactions without further purification.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for dPCR Experiments
| Reagent | Function | Considerations |
|---|---|---|
| Hot-Start DNA Polymerase | Enables specific amplification; reduces primer-dimer formation | Choose antibody-based, affibody, or chemically modified versions [66] |
| Hydrolysis Probes (TaqMan) | Sequence-specific detection with fluorophore-quencher pairs | Ensure minimal spectral overlap in multiplex applications [63] [67] |
| DNA-Binding Dyes (EvaGreen) | Binds dsDNA nonspecifically; cost-effective for single-plex | Not recommended for multiplexing without additional discrimination methods [63] |
| Nuclease-Free TE Buffer | Primer and probe resuspension; maintains oligo stability | Use pH 8.0 for most probes; pH 7.0 for Cy5 and Cy5.5 probes [63] |
| Restriction Enzymes | Digests complex DNA structures for even partitioning | Select enzymes that don't cut within amplicon sequence [63] |
| PCR Additives (DMSO, GC Enhancers) | Improves amplification of difficult templates (GC-rich regions) | Optimize concentration as they can affect primer Tm [66] [13] |
Workflow and Schematic Diagrams
dPCR Workflow with Primer-Dimer Mitigation
This workflow illustrates the complete dPCR process with integrated strategies to minimize primer-dimer formation, including hot-start enzymes and optimized primer design.
dPCR Multiplex Detection Schemes
This diagram illustrates the two primary duplex dPCR configurations: competing duplex for SNP/variant detection and non-competing duplex for copy number variation studies.
Digital PCR (dPCR) is a powerful technique for the absolute quantification of nucleic acids. The core principle involves partitioning a PCR reaction into thousands of individual reactions, so that a single DNA molecule can be amplified and detected in a binary manner (positive or negative). The absolute quantity of the target is then determined using Poisson statistics [69]. The two main platforms discussed here, Droplet Digital PCR (ddPCR) and nanoplate-based dPCR (ndPCR), differ primarily in their method of partition creation.
Droplet Digital PCR (ddPCR) uses a water-oil emulsion to generate tens of thousands of nanoliter-sized droplets, which act as the individual reaction chambers [70] [69]. Nanoplate-based dPCR (ndPCR), such as the QIAGEN QIAcuity, distributes the reaction mix into a microfluidic plate containing a fixed array of nanoscale wells [70] [71]. This fundamental difference in partitioning technology influences their workflow, performance, and suitability for different laboratory environments.
FAQs & Troubleshooting Guides
FAQ 1: How do the workflows of ddPCR and ndPCR differ, and which is more suited for a high-throughput QC lab?
Answer: The workflows are fundamentally different. The ndPCR workflow is more integrated and streamlined, making it generally more suitable for quality control (QC) environments where time, reproducibility, and ease of use are critical [72].
The table below summarizes the core workflow differences:
| Workflow Step | Droplet Digital PCR (ddPCR) | Nanoplate dPCR (ndPCR) |
|---|---|---|
| Partitioning | Multiple instruments: separate droplet generator required [70]. | Single instrument: integrated partitioning [70] [64]. |
| Thermocycling | Requires a conventional thermocycler [70] [73]. | Integrated into the dPCR instrument [70] [64]. |
| Reading/Analysis | Requires a separate droplet reader [70]. | Integrated imaging and analysis [70] [64]. |
| Hands-on Time | High; multiple pipetting and transfer steps [70] [72]. | Low; minimal transfer, similar to qPCR [70] [71]. |
| Total Time | Can be 6-8 hours [72]. | Around 2 hours for a full run [70]. |
| Contamination Risk | Higher due to multiple open-tube steps [70]. | Lower as reactions are confined to a sealed plate [70]. |
Answer: In ddPCR, the "rain" effect—droplets with intermediate fluorescence that are difficult to classify as positive or negative—is a common issue. It can be caused by several factors [70]:
- Droplet Variability: Inherent variability in droplet size and shape can affect amplification efficiency and fluorescence [70].
- Damaged Droplets: Droplets can coalesce or shear during thermal cycling, leading to aberrant signals [70].
- Non-specific Amplification: Primer-dimers or off-target amplification can create intermediate fluorescence [70] [63].
ndPCR minimizes these issues by using fixed, uniform partitions. The size and volume of the nanowells are consistent, which enhances the robustness and reproducibility of the method by eliminating variability associated with droplet generation [70] [74].
FAQ 3: My target is a large, complex DNA template (e.g., genomic DNA). How should I prepare my sample for accurate dPCR quantification?
Answer: Accurate quantification in dPCR relies on the random distribution of template molecules. Large or complex DNA structures can lead to uneven partitioning and over-quantification. For templates like high-molecular-weight genomic DNA, linked gene copies, or supercoiled plasmids, restriction enzyme digestion is highly recommended prior to the dPCR assay [63].
- Purpose: Digestion reduces viscosity, prevents multiple linked copies from being counted as a single molecule, and linearizes plasmids to improve primer/probe accessibility [63].
- Enzyme Selection: The restriction enzyme must not cut within the amplicon sequence itself [63]. Research indicates that the choice of enzyme can impact precision; one study found that HaeIII provided higher precision compared to EcoRI, especially for the QX200 ddPCR system [69].
FAQ 4: I am detecting low-abundance targets. How do the sensitivity of these platforms compare?
Answer: Both platforms offer high sensitivity, often superior to qPCR. Direct comparative studies have shown that both can detect very low copy numbers.
| Platform | Example Limit of Detection (LOD) | Comparison Context |
|---|---|---|
| ddPCR (Bio-Rad QX200) | 0.17 copies/µL input [69] | Detection of synthetic oligonucleotides and protist DNA. |
| ndPCR (QIAGEN QIAcuity) | 0.39 copies/µL input [69] | Detection of synthetic oligonucleotides and protist DNA. |
| ddPCR | 0.26 copies/µL [73] | Detection of Porcine Epidemic Diarrhea Virus (PEDV). |
| ndPCR | 1.83 copies/µL [71] | Detection of Canine Respiratory Coronavirus (CRCoV). |
One study concluded that both the QX200 and QIAcuity One demonstrated similar detection and quantification limits and yielded high precision across most analyses [69].
Experimental Protocols for Platform Comparison
For a rigorous comparison between ddPCR and ndPCR platforms, the following methodological approach, adapted from recent studies, is recommended.
Protocol: Cross-Platform Evaluation of dPCR Performance
1. Sample Preparation
- Use a combination of synthetic oligonucleotides (for a defined standard) and DNA extracted from a biological source (e.g., cell lines) [69].
- For biological DNA, use a series of increasing cell numbers to create a linearity curve [69].
- Perform restriction digestion on a subset of the biological DNA samples with different enzymes (e.g., HaeIII and EcoRI) to evaluate the impact on gene copy number quantification and precision [69].
2. Assay Design and Optimization
- Use the same primer and probe sequences for both platforms to ensure a direct comparison [64].
- Optimize primer and probe concentrations for each platform. For ddPCR, a typical optimal ratio may be 300nM primer : 200nM probe [73]. For ndPCR, concentrations may be adapted directly from a robust qPCR assay [71].
- Determine the optimal annealing temperature for the assay using a temperature gradient [73].
3. dPCR Run and Data Analysis
- Run the same samples on both platforms in parallel. For example, use the Bio-Rad QX200 ddPCR system and the QIAGEN QIAcuity One ndPCR system [69] [64].
- Calculate the Limit of Detection (LOD) and Limit of Quantification (LOQ) for both platforms from the serial dilution data [69].
- Assess precision by calculating the Coefficient of Variation (%CV) across replicates for each sample and platform [69] [64].
- Evaluate accuracy by comparing the measured gene copy numbers against the expected values from the synthetic standard and the cell count series [69].
The workflow for this comparative experiment is summarized in the diagram below.
The Scientist's Toolkit: Essential Research Reagent Solutions
The following table details key reagents and materials critical for successful dPCR experiments, based on the cited protocols.
| Item | Function & Importance |
|---|---|
| Restriction Enzymes (e.g., HaeIII) | Fragments large DNA templates to ensure random distribution and accurate quantification. Critical for analyzing complex genomic DNA, linked gene copies, or supercoiled plasmids [63] [69]. |
| High-Purity Primers & Probes | Sequence-specific detection. Must be stored in TE buffer (pH 8.0; pH 7.0 for Cy5-labeled probes) to prevent degradation. Higher concentrations (e.g., 0.5-0.9 µM primers, 0.25 µM probe) often yield better fluorescence intensity in dPCR [63]. |
| dPCR Supermix | Provides the core components for PCR amplification (polymerase, dNTPs, buffer). Formulations are often platform-specific. Using a "hot-start" polymerase can help reduce primer-dimer formation [73] [34]. |
| Nuclease-Free Water | Used to reconstitute primers/probes and adjust reaction volume. Essential for avoiding RNase and DNase contamination that can degrade nucleic acids and reagents [73]. |
| No-Template Control (NTC) | A critical control containing all reaction components except the template DNA/RNA. Used to monitor for contamination and primer-dimer formation [63] [73]. |
The transfer of quantitative PCR (qPCR) methods to digital PCR (dPCR) represents a significant advancement in molecular detection, offering absolute quantification without standard curves and enhanced resilience to amplification inhibitors. This transition is particularly valuable within the broader context of reducing primer-dimer formation in PCR research, as the partitioning step in dPCR inherently mitigates the impact of such nonspecific amplification products that can compromise qPCR accuracy. This guide addresses the key procedural adaptations and verification protocols required for a successful transition, providing a foundational resource for researchers and drug development professionals.
Fundamental Concepts: How dPCR Differs from qPCR
Q: What is the core technological difference between qPCR and dPCR that improves performance?
A: The fundamental difference lies in sample partitioning. qPCR performs amplification in a single, bulk reaction, where the cycle threshold (Ct) is used to infer the initial template concentration relative to a standard curve. In contrast, dPCR divides the reaction mixture into thousands to millions of individual partitions, so that each contains either zero, one, or a few template molecules [75]. Following end-point PCR amplification, the partitions are analyzed as positive or negative, and the absolute initial template concentration is calculated directly using Poisson statistics [75]. This partitioning reduces competition from primer-dimers and other nonspecific products, as they are confined to a subset of reactions rather than dominating the entire bulk mixture [11].
The diagram below illustrates the core workflow and this key difference.
Core Protocol: Adapting a qPCR Assay for dPCR
Q: What are the specific steps required to adapt my existing qPCR primers and probe for use with a dPCR system?
A: The adaptation process involves optimizing both the reaction composition and the thermal cycling conditions. The following protocol is based on the Bio-Rad QX200 Droplet Digital PCR system, but the principles are applicable to other platforms [76].
Reaction Setup and Master Mix
A standard 20-25 μL reaction volume is typical. The master mix is prepared as follows [76]:
- ddPCR Supermix: 12.5 μL of a 2X concentrate. This contains the DNA polymerase, dNTPs, and an optimized buffer.
- Primers and Probe: A 20X primer/probe mix is prepared so that adding 1-2.25 μL to the reaction gives the final desired concentration. It is crucial to use the same stringent primer design rules as for qPCR to minimize primer-dimer potential: primers should be 15-30 bases, have a GC content of 40-60%, and avoid complementary regions at the 3' ends [20] [13].
- Template DNA: The amount required depends on target abundance. For single-copy genes, 100 ng of genomic DNA is a common starting point. The template should be high-quality and free of inhibitors. For complex genomes, restriction digestion of 1 μg DNA prior to dilution and setup is recommended to ensure efficient partitioning [76].
- Nuclease-free Water: Added to bring the reaction to the final volume.
Droplet Generation and Thermal Cycling
- Droplet Generation: The reaction mixture is loaded into a dedicated cartridge along with droplet generation oil. The cartridge is placed in a droplet generator, which partitions each sample into approximately 20,000 nanodroplets [76].
- Thermal Cycling: The droplets are transferred to a PCR plate, sealed, and placed in a thermal cycler. While conditions are similar to qPCR, optimization is often needed:
- Annealing Temperature: A temperature gradient (e.g., 55°C to 60°C) should be run for new assays to determine the optimal temperature for specificity [76].
- Ramp Rate: A slower ramp rate (e.g., 3°C/sec) is recommended to ensure temperature uniformity across all droplets [76].
- Cycle Number: Because dPCR is an end-point measurement, the number of cycles only needs to be sufficient to robustly distinguish positive from negative partitions; typically, 40 cycles is standard.
The table below summarizes the key experimental parameters to optimize during transfer.
Table 1: Key Parameters for qPCR to dPCR Method Transfer
| Parameter | qPCR Typical Use | dPCR Optimization & Consideration |
|---|---|---|
| Template Input | Relies on standard curve for quantification. | 100 ng gDNA for single-copy targets; may require reduction for high-copy targets to avoid saturation [76]. |
| Annealing Temperature | Optimized for efficiency and specificity. | Re-optimize using a gradient; can often be increased by 2-5°C for enhanced specificity [13]. |
| Primer/Probe Concentration | Typically 200-900 nM for primers, 100-250 nM for probes. | Final concentration is critical; must be optimized for the specific ddPCR mastermix used [77]. |
| Data Analysis | Cycle threshold (Ct) relative to a standard curve. | Absolute quantification (copies/μL) based on Poisson statistics from positive/negative droplet counts [75]. |
Performance Verification and Validation
Q: After adapting my assay, how do I verify that the dPCR method is performing accurately and robustly?
A: A systematic validation is required to confirm the method's performance characteristics. A multifactorial experimental design is recommended to demonstrate robustness [77].
Key Performance Metrics
- Limit of Detection (LOD) and Quantification (LOQ): The LOD is the lowest concentration at which the target can be reliably detected, while the LOQ is the lowest concentration that can be accurately quantified. For example, a validated HDV RT-dPCR assay achieved an LOD of 0.7 copies/mL and an LOQ of 10 copies/mL [75].
- Linearity and Dynamic Range: The method should be linear across the expected concentration range. This can be demonstrated by testing a series of serial dilutions of a reference material.
- Accuracy (Trueness) and Precision: Accuracy can be established by measuring a recognized international standard (e.g., the WHO HDV international standard) and calculating a conversion factor to International Units (IU/mL) [75]. Precision, both within-run and between-run, should be evaluated, with a standard deviation of ±1.12 log IU/mL reported in one study as acceptable for viral load detection [75].
- Robustness: The method should be tested for resilience to factors like different operators, reagent lots, and incubation times. Studies show that ddPCR is generally very robust, with factors like operator and primer/probe system having no relevant effect on quantification [77].
Table 2: Essential Reagents and Materials for dPCR Setup
| Item | Function | Example |
|---|---|---|
| ddPCR Supermix | Provides core PCR components (polymerase, dNTPs, buffer). Critical for accuracy; different mixes can yield different results [77]. | Bio-Rad ddPCR Supermix for Probes (no dUTP) [77]. |
| Droplet Generation Oil | Creates the water-in-oil emulsion for partitioning the reaction. | Bio-Rad Droplet Generation Oil [76]. |
| Digestion Restriction Enzymes | Used to digest genomic DNA to ensure free partitioning of target sequences and prevent bias from linked DNA fragments. | Various (e.g., HindIII, EcoRI) [76]. |
| Cartridges and Gaskets | Microfluidic consumables used to generate droplets in specific systems. | Bio-Rad 8-chamber cartridges and rubber gaskets [76]. |
| Pierceable Foil Heat Seal | Used to securely seal the PCR plate during thermal cycling, preventing cross-contamination and droplet loss. | Bio-Rad Pierceable Foil Heat Seal [76]. |
Troubleshooting Common Issues in dPCR
Q: What are some common problems encountered during the qPCR to dPCR transfer, and how can they be resolved?
A: The following table addresses frequent challenges and provides solutions based on technical guides and published literature.
Table 3: Troubleshooting Guide for dPCR Method Transfer
| Problem | Potential Cause | Recommended Solution |
|---|---|---|
| Low Number of Accepted Droplets | Cartridge or gasket failure; improper pipetting; clogged microfluidics. | Ensure proper loading of sample and oil; check that the gasket is correctly seated; clean droplet generator [76]. |
| Poor Resolution Between Positive and Negative Populations | Suboptimal annealing temperature; probe degradation; insufficient PCR efficiency. | Perform annealing temperature gradient; prepare fresh probe aliquots; check primer/probe design and concentration [13] [76]. |
| Inaccurate Quantification (Bias) | Template overloading; inadequate restriction digestion; suboptimal master mix. | Reduce template DNA input to prevent partition saturation; ensure complete restriction digestion of gDNA; test different master mixes [77] [76]. |
| High Background or Non-specific Amplification | Primer-dimer formation in partitions; low annealing temperature; contaminated reagents. | Re-design primers to avoid 3' complementarity; increase annealing temperature; use hot-start DNA polymerases; prepare fresh reagents [11] [13]. |
The following diagram outlines a logical workflow for diagnosing and resolving the most common dPCR issues.
This technical support guide provides comprehensive troubleshooting resources for validating reference genes in RT-qPCR experiments. Proper reference gene selection is crucial for obtaining accurate gene expression data, particularly when aiming to reduce primer-dimer formation and other amplification artifacts in PCR research. The following sections address common challenges and provide optimized protocols to ensure experimental reliability.
FAQs: Reference Gene Selection and Validation
1. Why is reference gene validation critical for accurate RT-qPCR results?
Reference gene validation is essential because inappropriate reference genes can lead to misinterpretation of gene expression data. Studies have demonstrated that using different reference genes can yield surprisingly different results [78]. According to MIQE guidelines, normalizing RT-qPCR data against a single reference gene is no longer acceptable, and reference genes must be validated for each specific experimental condition [79]. For example, research on Pseudomonas aeruginosa L10 under n-hexadecane stress revealed that different algorithms identified different genes as most stable, highlighting the need for comprehensive validation [80].
2. How does primer-dimer formation affect reference gene validation?
Primer-dimer formation compromises RT-qPCR accuracy by competing with the target amplification for reagents, potentially leading to false quantification of reference gene expression [11]. Primer-dimers are short, unintended DNA fragments that form when primers anneal to each other instead of the target DNA, resulting in nonspecific amplification that can significantly hinder PCR efficiency and accuracy [2]. This is particularly problematic in reference gene validation where precise quantification is essential.
3. What are the most common causes of primer-dimer formation?
- Inadequate primer design: Primers with self-complementary regions or complementary 3' ends [5]
- Low annealing temperatures: Promotes nonspecific binding [13]
- High primer concentration: Increases likelihood of primer-primer interactions [2]
- Poor quality reagents: Contaminants can promote dimer formation [11]
- Excessive PCR cycles: Allows accumulation of primer-dimers after target amplification plateaus [5]
- Improper reaction setup: Leaving reactions at room temperature before thermal cycling [2]
4. Which algorithms are recommended for assessing reference gene stability?
Multiple algorithms should be used for comprehensive reference gene validation [79]. The most widely accepted tools include:
- geNorm: Measures expression stability by calculating stability value (M) [81]
- NormFinder: Uses model-based approach considering intra- and inter-group variation [79]
- BestKeeper: Evaluates stability through correlation analysis of Cq values [80]
- RefFinder: Provides comprehensive ranking by integrating multiple algorithms [81]
Troubleshooting Guides
Guide 1: Preventing Primer-Dimer Formation in Reference Gene Assays
| Problem | Possible Causes | Solutions |
|---|---|---|
| Primer-dimer in no-template control | Primer complementarity, low annealing temperature, excessive cycles | Redesign primers with less 3' complementarity, increase annealing temperature, optimize cycle number [5] |
| Smear or multiple bands on gel | Non-specific binding, contaminated reagents, insufficient annealing temperature | Use hot-start DNA polymerase, optimize Mg2+ concentration, increase annealing temperature gradient [13] |
| Inconsistent Cq values between replicates | Poor RNA quality, pipetting errors, insufficient mixing | Check RNA integrity, use proper pipetting technique, mix reagents thoroughly [13] |
| High background signal in qPCR | Excessive primer concentration, probe degradation, contaminated cDNA | Lower primer concentration, protect probes from light, use UV-irradiated workspace [45] |
Guide 2: Reference Gene Validation Workflow Issues
| Problem | Possible Causes | Solutions |
|---|---|---|
| Variable reference gene expression across conditions | True biological variation, inappropriate gene selection | Test multiple candidates, use algorithms to identify most stable genes [78] |
| Discrepant stability rankings between algorithms | Different statistical approaches, sample heterogeneity | Use comprehensive approach (RefFinder), ensure adequate sample size [80] |
| Poor amplification efficiency | Primer design issues, PCR inhibitors, suboptimal conditions | Check primer specificity, purify template, optimize buffer conditions [13] |
| Inconsistent results between technical replicates | Equipment calibration issues, reaction setup errors | Calibrate pipettes, use master mixes, verify thermal cycler performance [79] |
Experimental Protocols
Protocol 1: Comprehensive Reference Gene Validation
Materials:
- RNA samples from all experimental conditions
- cDNA synthesis kit
- qPCR reagents and equipment
- Primer sets for candidate reference genes
Procedure:
- Select Candidate Reference Genes: Choose 3-8 genes based on literature and preliminary data [79]. Common candidates include:
- act (actin), atub (α-tubulin), btub (β-tubulin)
- ef1 (elongation factor 1-α), ef2 (elongation factor 2)
- rpsL (ribosomal protein), gyrA (DNA gyrase) [80]
RNA Extraction and Quality Control:
- Extract RNA using standardized methods
- Assess purity (A260/280 ratio of 1.9-2.1) [79]
- Verify integrity via gel electrophoresis (sharp 18S and 28S rRNA bands)
cDNA Synthesis:
- Use consistent input RNA amounts across samples
- Include no-reverse transcriptase controls
qPCR Amplification:
- Run samples in technical triplicates
- Include no-template controls for each primer set
- Use the following cycling conditions:
- Initial denaturation: 95°C for 3-5 minutes
- 40 cycles of: 95°C for 15-30s, annealing at optimized temperature for 30s, 72°C for 30s
- Final extension: 72°C for 5-10 minutes [13]
Data Analysis:
- Calculate amplification efficiencies (90-110% acceptable)
- Export Cq values for stability analysis
- Analyze with geNorm, NormFinder, BestKeeper, and RefFinder
- Select the most stable reference genes (M value < 0.5 in geNorm) [79]
Protocol 2: Primer-Dimer Minimization for Reference Gene Assays
Materials:
- HPLC-purified primers
- Hot-start DNA polymerase
- Gradient thermal cycler
Procedure:
- Primer Design Optimization:
Reaction Setup:
Thermal Cycling Optimization:
Reference Gene Stability: Comparative Data
Table 1: Reference Gene Stability Across Experimental Conditions
| Experimental System | Most Stable Reference Genes | Least Stable Reference Genes | Validation Tools Used | Reference |
|---|---|---|---|---|
| E. coli under antimicrobial blue light | ihfB, cysG, uidA, gyrA | Not specified | GeNorm, NormFinder, BestKeeper, RefFinder | [81] |
| Phytophthora capsici during infection | ef1, ws21, ubc | atub, ef2 | GeNorm, NormFinder, BestKeeper, ΔCt method | [79] |
| P. aeruginosa under n-hexadecane stress | nadB, anr | tipA | GeNorm, NormFinder, BestKeeper, RefFinder | [80] |
| Four grasshopper species | Species and tissue-dependent | Species and tissue-dependent | GeNorm, NormFinder, BestKeeper, ΔCt method | [78] |
Table 2: Troubleshooting Primer-Dimer Formation
| Factor | Optimal Conditions | Common Pitfalls | Impact on Primer-Dimer |
|---|---|---|---|
| Primer Design | 18-24 nt; 40-60% GC; Tm 54-65°C | 3' complementarity; high GC clamps | High impact [7] |
| Annealing Temperature | 3-5°C below Tm; optimized via gradient | Too low temperature | High impact [13] |
| Primer Concentration | 0.1-1.0 μM; optimized for each assay | Excessive concentration | High impact [2] |
| DNA Polymerase | Hot-start varieties | Standard polymerase added too early | Moderate impact [5] |
| Cycle Number | 25-40 cycles | Excessive cycles | Moderate impact [13] |
| Template Quality | High purity, no inhibitors | Contaminated or degraded | Variable impact [13] |
Research Reagent Solutions
Table 3: Essential Materials for Reference Gene Validation
| Reagent | Function | Recommended Specifications |
|---|---|---|
| Hot-Start DNA Polymerase | Amplification with reduced pre-cycling activity | High fidelity, proofreading capability for reference gene applications [13] |
| RNA Extraction Kit | High-quality RNA isolation | Consistent yield, genomic DNA removal, maintained RNA integrity [79] |
| Reverse Transcriptase | cDNA synthesis from RNA templates | High efficiency, minimal RNase contamination [80] |
| qPCR Master Mix | Quantitative detection | Consistent performance, low background, compatible with multiplexing [81] |
| Primer Design Software | In silico primer optimization | Self-complementarity analysis, specificity checking, Tm calculation [7] |
| Nucleic Acid Quantification | Sample quality assessment | Spectrophotometric or fluorometric capability, low sample requirement [79] |
Workflow Diagrams
Reference Gene Validation and Primer-Dimer Prevention Workflow
Multi-Algorithm Approach to Reference Gene Validation
Conclusion
Effective primer-dimer management requires an integrated approach combining thoughtful primer design, optimized reaction conditions, systematic troubleshooting, and rigorous validation. The strategic implementation of hot-start polymerases, precise temperature control, and appropriate primer modifications significantly enhances PCR specificity and reliability. As molecular diagnostics and precision medicine advance, emerging technologies like digital PCR offer powerful alternatives for absolute quantification with reduced dimer susceptibility. Future directions will likely incorporate artificial intelligence for predictive primer design and novel polymerase engineering for enhanced specificity, ultimately enabling more accurate genetic analysis across biomedical research, therapeutic development, and clinical applications. By adopting the comprehensive strategies outlined, researchers can transform primer-dimer challenges into opportunities for assay optimization and result verification.
References
- 1. Primer dimer [https://en.wikipedia.org/wiki/Primer_dimer]
- 2. Troubleshooting primer dimer in PCR [https://www.minipcr.com/primer-dimer-pcr/]
- 3. Primer Dimer - an overview [https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/primer-dimer]
- 4. What are Primer Dimers? - A Beginner's Guide [https://geneticeducation.co.in/what-are-primer-dimers-a-beginners-guide/]
- 5. PCR Troubleshooting 103: How to Address Primer-Dimers [https://geneticeducation.co.in/pcr-troubleshooting-103-how-to-address-primer-dimers/]
- 6. Eliminating primer dimers and improving SNP detection ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7200914/]
- 7. Primer design guide - 5 tips for best PCR results [https://the-dna-universe.com/2022/09/05/primer-design-guide-the-top-5-factors-to-consider-for-optimum-performance/]
- 8. How can I avoid primer-dimer formation during PCR ... [https://www.qiagen.com/us/resources/faq/544]
- 9. Optimizing PCR: Proven Tips and Troubleshooting Tricks [https://www.the-scientist.com/optimizing-pcr-proven-tips-and-troubleshooting-tricks-71660]
- 10. The Pain of Primer Dimer [https://kilobaser.com/post/the-pain-of-primer-dimer]
- 11. PCR Optimization: Strategies To Minimize Primer Dimer [https://genemod.net/blog/pcr-optimization]
- 12. Primer Dimer - an overview [https://www.sciencedirect.com/topics/neuroscience/primer-dimer]
- 13. PCR Troubleshooting Guide | Thermo Fisher Scientific - US [https://www.thermofisher.com/us/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/pcr-troubleshooting.html]
- 14. Impact of Primer Dimers and Self-Amplifying Hairpins on ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5922443/]
- 15. Optimizing PCR Reaction Conditions for High Fidelity and ... [https://www.labx.com/resources/optimizing-pcr-reaction-conditions-for-high-fidelity-and-yield/5579]
- 16. PCR Optimization: Factors Affecting Reaction Specificity ... [https://www.mybiosource.com/learn/pcr-optimization-factors-affecting-reaction-specificity-and-efficien/]
- 17. Error-suppression mechanism of PCR by blocker strands - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10111364/]
- 18. Preventing False Positive and False Negative PCR Results [https://www.clinicallab.com/preventing-false-positive-and-false-negative-pcr-results-26261]
- 19. DNA Amplification | No Complementarily to other primers [https://nij.ojp.gov/nij-hosted-online-training-courses/dna-amplification/overview/primer-design/no-complementarily-other-primers]
- 20. Polymerase Chain Reaction: Basic Protocol Plus ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4846334/]
- 21. PCR Troubleshooting Guide [https://www.neb.com/en-us/tools-and-resources/troubleshooting-guides/pcr-troubleshooting-guide?srsltid=AfmBOoohSqvxg8FZTXjKvG5BvWwzgnqdvANa6IDyNZDbSWEY-i-SR9sU]
- 22. PCR Primer Design Tips - Behind the Bench [https://www.thermofisher.com/blog/behindthebench/pcr-primer-design-tips/]
- 23. Primer designing tool - NCBI - NIH [https://www.ncbi.nlm.nih.gov/tools/primer-blast/]
- 24. Four tips for PCR amplification of GC-rich sequences [https://www.neb.com/en-us/nebinspired-blog/four-tips-for-pcr-amplification-of-gc-rich-sequences?srsltid=AfmBOoqH-JvfaEXPKcDfkxM7-1q1JNkwUu8RL-jHj15DuqdHyowv4AH1]
- 25. Tips for Designing PCR Primers [https://blog.edvotek.com/2025/08/14/tips-for-designing-pcr-primers/]
- 26. PrimerChecker [https://primerchecker.okstate.edu/]
- 27. Multiple Primer Analyzer | Thermo Fisher Scientific - US [https://www.thermofisher.com/us/en/home/brands/thermo-scientific/molecular-biology/molecular-biology-learning-center/molecular-biology-resource-library/thermo-scientific-web-tools/multiple-primer-analyzer.html]
- 28. Self-avoiding molecular recognition systems (SAMRS ... [https://www.biosyn.com/tew/Members-of-the-%E2%80%9CSelf-Avoiding-Molecular-Recognition-Systems%E2%80%9D-or-SAMRS-bind-to-natural-DNA-but-not-to-other-SAMRS-analogs.aspx]
- 29. Artificial Genetic Systems. Self-Avoiding DNA in PCR and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6027612/]
- 30. Locked Nucleic Acids (LNAs) in PCR Assays [https://kcasbio.com/blogs/locked-nucleic-acids-lnas-in-pcr-assays-enhanced-sensitivity-and-specificity/]
- 31. Locked nucleic acids in PCR primers increase sensitivity ... [https://pubmed.ncbi.nlm.nih.gov/18164179/]
- 32. Utilization of locked nucleic acids in PCR primers and ... [https://www.npdn.org/newsletter/2025/05/utilization_locked_nucleic_acids_pcr_primers_and_probes_enhance_specificity]
- 33. 减少PCR引物浓度以避免二聚体的方法小结 [https://bhsy-e.biomart.cn/news/3238726.htm]
- 34. PCR Troubleshooting: Common Problems and Solutions [https://www.mybiosource.com/learn/pcr-troubleshooting-common-problems-and-solutions/]
- 35. PCR Troubleshooting Guide [https://www.neb.com/en-us/tools-and-resources/troubleshooting-guides/pcr-troubleshooting-guide?srsltid=AfmBOopbqDT56hiZH6bn6YTNnyfYE70t-SM59qCAEvuFu161Tu7ZhHkr]
- 36. Prediction of PCR amplification from primer and template ... [https://www.nature.com/articles/s41598-021-86357-1]
- 37. Hot Start PCR with heat-activatable primers [https://pmc.ncbi.nlm.nih.gov/articles/PMC2582603/]
- 38. Optimizing your PCR [https://www.takarabio.com/learning-centers/pcr/faq/optimization?srsltid=AfmBOopyiUFdFMfSEVQ9vBZDeaImdMOGqEFksNW5L0MdDInEB66PNO9a]
- 39. How to Simplify PCR Optimization Steps for Primer Annealing [https://www.thermofisher.com/us/en/home/brands/thermo-scientific/molecular-biology/molecular-biology-learning-center/molecular-biology-resource-library/spotlight-articles/pcr-annealing-optimization-universal-annealing.html]
- 40. How is Hot-Start Technology Beneficial For Your PCR [https://www.thermofisher.com/us/en/home/brands/thermo-scientific/molecular-biology/molecular-biology-learning-center/molecular-biology-resource-library/spotlight-articles/hot-start-technology-benefits-PCR.html]
- 41. Amplification of the No Template Control (NTC) [https://www.thermofisher.com/us/en/home/life-science/pcr/real-time-pcr/real-time-pcr-learning-center/real-time-pcr-basics/real-time-pcr-troubleshooting-tool/gene-expression-quantitation-troubleshooting/amplification-no-template-control.html]
- 42. Why Your Negative Control Has a Band (And How to Fix It) [https://www.clyte.tech/post/troubleshooting-pcr-why-your-negative-control-has-a-band-and-how-to-fix-it?srsltid=AfmBOooIvTTdUXs_UpOiA8YZ2EfE7iD4x6sFFpydx5pptFTjCUHgugNL]
- 43. Prevention of pre-PCR mis-priming and primer ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC312262/]
- 44. Nucleic Acid Gel Electrophoresis Troubleshooting [https://www.thermofisher.com/us/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/na-electrophoresis-education/na-electrophoresis-troubleshooting.html]
- 45. Optimization of primer sets and detection protocols for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7295692/]
- 46. PCR Setup—Six Critical Components to Consider [https://www.thermofisher.com/us/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/pcr-component-considerations.html]
- 47. Guidelines for PCR Optimization with Taq DNA Polymerase [https://www.neb.com/en-us/tools-and-resources/usage-guidelines/guidelines-for-pcr-optimization-with-taq-dna-polymerase?srsltid=AfmBOop1MJ4OdOLeeZpgwqFK4MfWDpWpKv_fA2VlJJ3Z0yj9hSMOL_VT]
- 48. DNA Polymerase—Four Key Characteristics for PCR [https://www.thermofisher.com/us/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/dna-polymerase-characteristics.html]
- 49. Enhancement of DNA amplification efficiency by using ... [https://www.bocsci.com/blog/enhancement-of-dna-amplification-efficiency-by-using-pcr-additives/?srsltid=AfmBOoojqgoRozAtnovSPZkuTkojTNC52MK0mGGknXXl8_2HC5RpjYX7]
- 50. Types, mechanisms, and applications in long-range PCR [https://www.sciencedirect.com/science/article/abs/pii/S0300908422000499]
- 51. Betaine, Dimethyl Sulfoxide, and 7-Deaza-dGTP, a Powerful ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1876170/]
- 52. Bovine serum albumin further enhances the effects of organic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3466135/]
- 53. Betaine, dimethyl sulfoxide, and 7-deaza-dGTP, a powerful ... [https://www.semanticscholar.org/paper/Betaine%2C-dimethyl-sulfoxide%2C-and-7-deaza-dGTP%2C-a-of-Musso-Bocciardi/aeb13a3833415e347b68820ee0d9a60b1bd35a31]
- 54. Fundamentals of Sequencing of Difficult Templates—An ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2291785/]
- 55. How to Amplify Difficult PCR Substrates [https://bitesizebio.com/9331/tips-for-amplifying-difficult-pcr-substrates/]
- 56. PCR Problems? Try an Additive [https://bitesizebio.com/24/pcr-problems-try-an-additive/]
- 57. Guidelines for PCR Optimization with Thermophilic DNA ... [https://www.neb.com/en-us/tools-and-resources/usage-guidelines/guidelines-for-pcr-optimization-with-thermophilic-dna-polymerases?srsltid=AfmBOorb517fxeErevRzXR_pF5IKEVyTUdb159kLGU037kt1s2WggClM]
- 58. PCR Cycling Parameters—Six Key Considerations for ... [https://www.thermofisher.com/us/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/pcr-cycling-considerations.html]
- 59. Troubleshooting your PCR [https://www.takarabio.com/learning-centers/pcr/faq/troubleshooting?srsltid=AfmBOopF4YWCUR80VlGf0FDAerPO3ohoYLM_KvxpvU4Q8Xadxr09WNqW]
- 60. - RNA / BOC Sciences Primer Dimers [https://rna.bocsci.com/support/primer-dimers-a-comprehensive-guide.html]
- 61. Core vs. Full Process Validation: Choosing the Right Path ... [https://www.eurofins-viracorbiopharma.com/resource-library/posts/2025/august/blog-core-vs-full-process-validation-choosing-the-right-path-for-pcr-assay-confidence/]
- 62. Fundamentals of multiplexing with digital PCR - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5154634/]
- 63. dPCR assay development | Digital PCR troubleshooting [https://www.qiagen.com/us/applications/digital-pcr/beginners/dpcr-guide/setup-and-troubleshooting]
- 64. Comparison of two digital PCR platforms for quantification ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12622311/]
- 65. PCR Troubleshooting Guide [https://www.neb.com/en-us/tools-and-resources/troubleshooting-guides/pcr-troubleshooting-guide?srsltid=AfmBOorPOsDXRc1sZY77gNSE_L1ZafXry2BNY6aKmpLIUg5gqGx7Agmw]
- 66. PCR Methods - Top Ten Strategies [https://www.thermofisher.com/us/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/pcr-methods.html]
- 67. Evolution of Multiplex PCR: Benefits, Challenges & Key ... [https://www.eurogentec.com/en/news/111_evolution-of-multiplex-pcr-benefits-challenges-key-applications]
- 68. Digital PCR Support—Troubleshooting [https://www.thermofisher.com/us/en/home/technical-resources/technical-reference-library/real-time-digital-PCR-applications-support-center/digital-pcr-support/digital-pcr-support-troubleshooting.html]
- 69. Comparing the precision of two digital PCR applications for ... [https://www.nature.com/articles/s41598-025-13143-8]
- 70. Comparison of digital PCR methods [https://www.qiagen.com/us/knowledge-and-support/knowledge-hub/bench-guide/pcr/digital-pcr/comparison-of-digital-pcr-methods]
- 71. Development and validation of nanoplate-based RT-dPCR ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12085003/]
- 72. dPCR vs. ddPCR: Why Precision Matters in Cell and Gene ... [https://www.roslinct.com/insights/dpcr-vs-ddpcr/]
- 73. Development of a droplet digital PCR for detection and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7438641/]
- 74. More less-frequently asked questions on dPCR [https://www.qiagen.com/us/knowledge-and-support/knowledge-hub/science-matters/pcr-solutions/more-less-frequently-asked-questions-on-dpcr]
- 75. Droplet Digital PCR: A Powerful Tool for Accurate ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12080313/]
- 76. dPCR Protocol [https://www.sigmaaldrich.com/US/en/technical-documents/protocol/genomics/qpcr/dpcr?srsltid=AfmBOoqkrFRiB0bJ2f4TRlVKbhiqBJj60gWaPrmAihStH7X9R-EDm_JC]
- 77. In-house validation of a droplet digital PCR system using ... [https://www.sciencedirect.com/science/article/pii/S0003267025005665]
- 78. the importance of testing reference gene stability in RT-qPCR ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7409783/]
- 79. Validation of reference gene stability for normalization ... [https://www.nature.com/articles/s41598-024-58139-y]
- 80. Identification and validation of reference genes for qPCR of ... [https://bmcmicrobiol.biomedcentral.com/articles/10.1186/s12866-025-04112-2]
- 81. Identification and validation of reference genes for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11743365/]